The srpAnalytics modeling pipeline for the Superfund Research Program Analytics Portal.
Filter results
Software Type
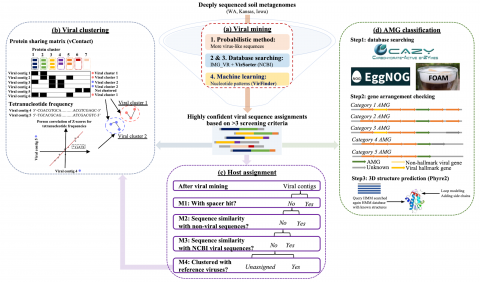
Code pertaining to the Soil Microbiome SFA Project publication data visualizations 'DNA viral diversity, abundance and functional potential vary across grassland soils with a range of historical moisture regimes' for processing publication data downloads.
Category
LIQUID Software Overview LIQUID provides users with the capability to process high throughput data and contains a customizable target library and scoring model per project needs. The graphical user interface provides visualization of multiple lines of spectral evidence for each lipid identification...
Category
pmartR Software Overview The pmartR package provides a single software tool for QC (filtering and normalization), exploratory data analysis (EDA), and statistical analysis (robust to missing data) and includes numerous visualization capabilities of mass spectrometry (MS) omics data (proteomic...
The following R source code was used for plotting figures of the viral communities detected from three grasslands soil metagenomes with a historical precipitation gradient ( WA-TmG.2.0 , KS-TmG.2.0 , IA-TmG.2.0 ) from project publication 'DNA viral diversity, abundance and functional potential vary...
Category
Fusarium sp. DS682 Proteogenomics Statistical Data Analysis of SFA dataset download: 10.25584/KSOmicsFspDS682/1766303 . GitHub Repository Source: https://github.com/lmbramer/Fusarium-sp.-DS-682-Proteogenomics MaxQuant Export Files (txt) Trelliscope Boxplots (jsonp) Fusarium Report (.Rmd, html)...
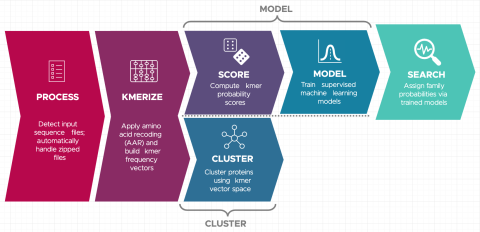
Last updated on 2023-02-23T19:37:46+00:00 by LN Anderson Snekmer: A scalable pipeline for protein sequence fingerprinting using amino acid recoding (AAR) Snekmer is a software package designed to reduce the representation of protein sequences by combining amino acid reduction (AAR) with the kmer...