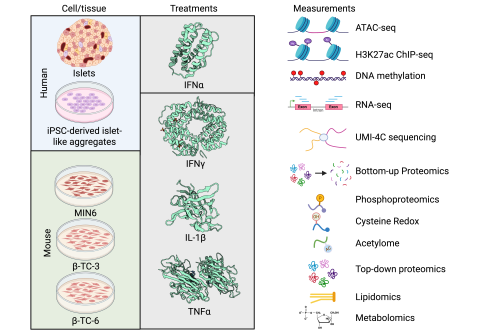
This data package consists of data from MIN6 cells (n=4) treated with IL-1β + IFNγ + TNFα and RNAi targeting Poly(ADP)ribosyl polymerase 12 ( Parp12 ) for 24 h and submitted to proteomic analyses ( Table 1 ). PARP12 is an interferon-induced protein known to target viral RNA to stress granules. It...
Filter results
Category
- Scientific Discovery (237)
- Biology (159)
- Earth System Science (107)
- Human Health (74)
- Integrative Omics (73)
- Microbiome Science (22)
- National Security (22)
- Computing & Analytics (12)
- Energy Resiliency (9)
- Chemical & Biological Signatures Science (8)
- Weapons of Mass Effect (8)
- Computational Research (6)
- Visual Analytics (6)
- Atmospheric Science (5)
- Data Analytics & Machine Learning (5)
- Materials Science (4)
- Renewable Energy (4)
- Coastal Science (3)
- Computational Mathematics & Statistics (3)
- Data Analytics & Machine Learning (3)
- Ecosystem Science (3)
- Bioenergy Technologies (2)
- Cybersecurity (2)
- Distribution (2)
- Electric Grid Modernization (2)
- Energy Storage (2)
- Grid Cybersecurity (2)
- Plant Science (2)
- Solar Energy (2)
- Subsurface Science (2)
- Transportation (2)
- Chemistry (1)
- Computational Mathematics & Statistics (1)
- Energy Efficiency (1)
- Grid Analytics (1)
- High-Performance Computing (1)
- Terrestrial Aquatics (1)
- Wind Energy (1)
Dataset Type
- Omics (121)
- Proteomics (62)
- Transcriptomics (55)
- Sequencing (53)
- Metabolomics (35)
- Lipidomics (31)
- Simulation Results (24)
- Amplicon (16S, ITS) (16)
- RNA-Seq (mRNA,miRNA) (11)
- Metagenomics (9)
- Microarray (9)
- Computed Analysis (8)
- Genomics (7)
- Microscopy (6)
- ChIP-seq (4)
- Electron Microscopy (3)
- Geospacial (3)
- Mass Spectra (2)
- Phenomics (2)
- Staff (2)
- Cybersecurity (1)
- Electromagnetic Spectrum (1)
- Raw Image (1)
- Whole genome sequencing (WGS) (1)
Tags
- Virology (61)
- Immune Response (46)
- Time Sampled Measurement Datasets (46)
- Differential Expression Analysis (41)
- Gene expression profile data (40)
- Homo sapiens (37)
- Mass spectrometry data (25)
- Multi-Omics (24)
- Predictive Phenomics (17)
- MERS-CoV (16)
- Mus musculus (16)
- Soil Microbiology (15)
- Viruses (14)
- Health (13)
- Virus (13)
- West Nile virus (11)
- Mass Spectrometry (10)
- Proteomics (10)
- sequencing (10)
- Synthetic (10)
- Influenza A (9)
- Ebola (7)
- Metagenomics (7)
- S. elongatus PCC 7942 (7)
- Resource Metadata (6)
- Tandem Mass Tag (6)
- Global Analysis (5)
- HCoV-299E (5)
- Microarray (5)
- Post-Translational Modifications (5)
This data package consists of data from parallel experiments compared to Pck001 but performed with EndoC-βH1 cells instead of human islets. Cells were treated with IFNγ + IL-1β for 48 h and submitted to RNA-seq (n=5), ATAC-seq (n=5), DNA methylation (n=5), and H3K27Ac ChIP-seq (n=4). In addition...
Category
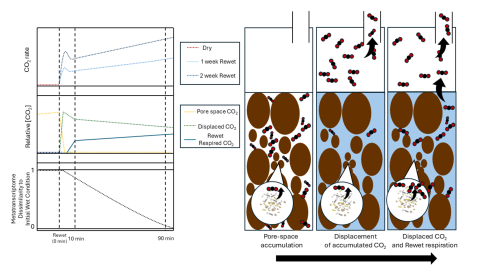
Please cite as : Mary S. Lipton, William C. Nelson, Hugh McCullough, Montana L. Smith, Sheryl L. Bell, Lisa M. Bramer, Hyun-Seob Song, Kirsten Hofmockel, 2025. Moisture Metaphenome Incubation Analysis Results . [Data Set] PNNL DataHub. https://doi.org/10.25584/3007775 This data is published under a...
RhizoGrid Indexed Sorghum Rhizosphere Multi-Omics Identification of spatially resolved biomarkers of drought in Sorghum bicolor rhizosphere molecular-microbe interactions using a novel root cartography "RhizoGrid" system for sampling plants under drought and control conditions across 10 equally...
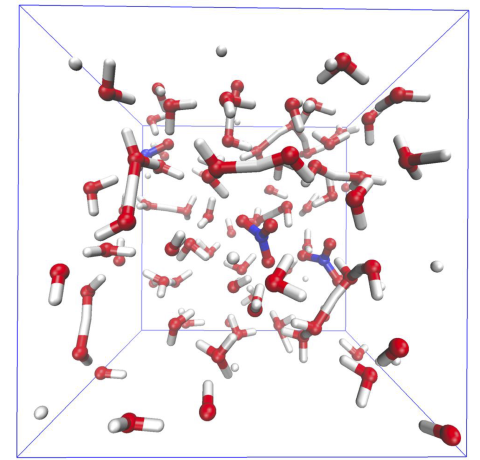
This dataset was generated using an iterative active learning strategy with the ArcaNN software package ( https://github.com/arcann-chem/arcann_training ) to train machine-learning interatomic potentials (MLIPs) for aqueous nitric acid. Each active-learning cycle consisted of three stages: (1)...
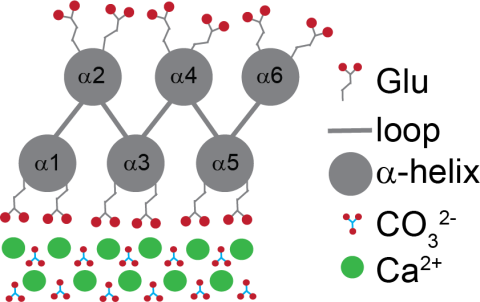
Biomineralization is the process by which organisms use biomolecules to produce hierarchically structured organic-inorganic composites. Using biology as inspiration, a protein construct (FD31) was previously designed to accelerate formation of nano-calcite with an unconventional {110} face. To...
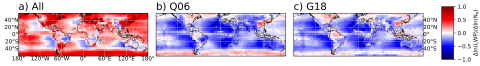
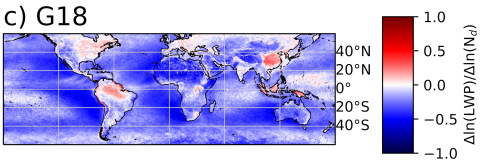
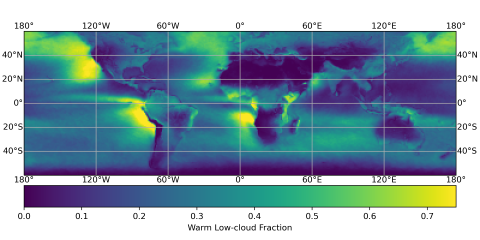
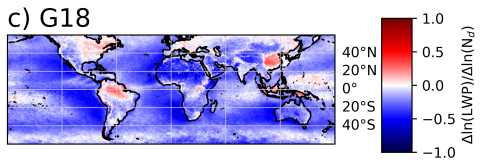
The SATELLITE_EAGLES_PNNL NetCDF dataset contains a suite of satellite- and reanalysis-derived atmospheric and surface parameters on a regular latitude–longitude grid. The dataset includes core geophysical fields such as land fraction, aerosol optical depth at multiple wavelengths (465, 550, 667...
The SATELLITE_EAGLES_PNNL NetCDF dataset contains a suite of satellite- and reanalysis-derived atmospheric and surface parameters on a regular 18 × 36 latitude–longitude grid. The dataset includes core geophysical fields such as land fraction, aerosol optical depth at multiple wavelengths (465, 550...
The SATELLITE_EAGLES_PNNL NetCDF dataset contains a suite of satellite- and reanalysis-derived atmospheric and surface parameters on a regular latitude–longitude grid. The dataset includes core geophysical fields such as land fraction, aerosol optical depth at multiple wavelengths (465, 550, 667...
The SATELLITE_EAGLES_PNNL NetCDF dataset contains a suite of satellite- and reanalysis-derived atmospheric and surface parameters on a regular latitude–longitude grid for 6 different spatial resolutions (10, 5, 1, 0.5, 0.1, and 0.05 degree resolutions). The dataset includes core geophysical fields...
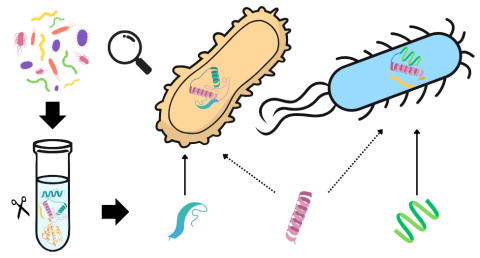
This data package contains processed LC-MS proteomics results and analysis scripts associated with the paper "Assessing Degenerate Peptide Resolution Methods using a Ground Truth Dataset". In this study, we designed an artificial microbial community to create a ground truth, simulated metaproteomic...
Category
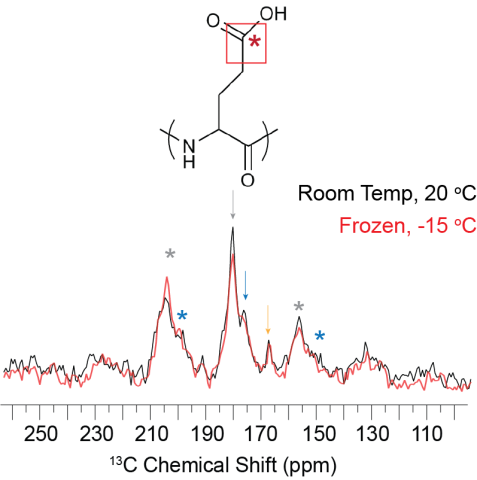
Room temperature and frozen CP-MAS spectra of FD31*-Edelta nucleated calcite showing the overall CSA pattern does not change as a function of temperature. The FD31* Glu sidechain carbonyl appears to be mobile on the ~500 microsecond timescale under both temperature conditions.
Raw NMR data for the three trials measuring the distance between 15N in the backbone of glutamic acid in FD31* to 13C in the calcite surface using REDOR NMR. Data collected on a 300 MHz instrument using a 5 mm HCN probe.
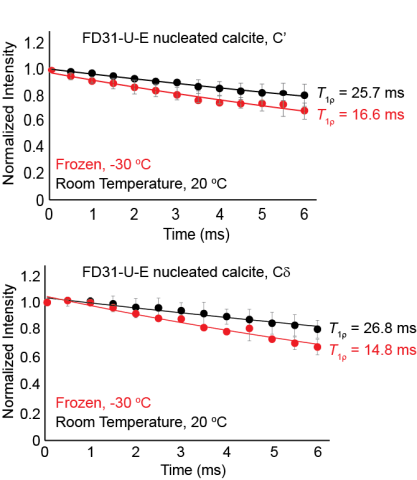
Raw data for all three trials used to calculate the T1rho value for FD31-U-E nucleated calcite at both room temperature and frozen. This data shows that both the FD31 backbone and sidechain carbonyls in glutamic are immobile in nucleated calcite on the millisecond timescale at both temperatures.
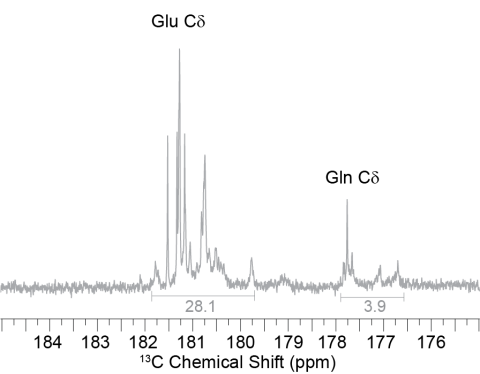
Solution NMR spectra of FD31*-Edelta dissolve in D2O, showing the distribution of peaks for glutamic acid (~180 ppm) and glutamine (~174 ppm), the latter of which was labeled via scrambling.