Description
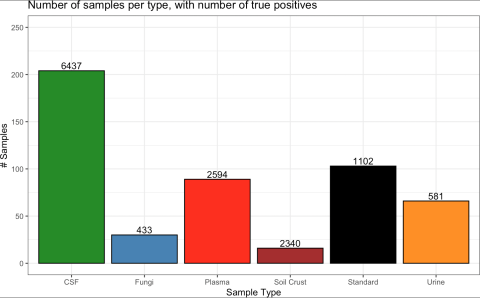
Data comprised of 204, 103, 89, 66, 30, and 16 gas chromatography mass spectrometry (GC-MS) runs from human cerebrospinal fluid (CSF), standards, human blood plasma, human urine, fungi species (A. niger, A. nidulans, T. reesei), and soil crust, respectively Across 4,523,251 candidate matches, 13,487 were hand-verified as true positives and 610,403 were determined to be true negatives. The remaining candidate matches were considered “unknown” due to the high presence of potential matches in complex samples where all possible compounds are not fully characterized.
Query spectra were converted from Agilent .D files to .cdf and matched to our reference database with CoreMS (https://zenodo.org/record/4641553#.Y4f75-zMKFA). Every possible metabolite within RI windows of size -/+35 RI, called an “RI bin”, was returned and subsequently hand-verified to determine true positives.
Sample Preparation and Data Acquisition. Standard metabolites were purchased from Sigma Aldrich. Dried metabolite standards were derivatized using a modified version of the FiehnLib protocol, as was described in Kind et al. 2009. Briefly, samples undergo methoximation to protect carbonyl groups and reduce tautomeric isomers. This was followed by silylation with N-methyl-N-trimethylsilyltrifluoroacetamide and 1% trimethylchlorosilane (MSTFA) to derivatize hydroxy and amine groups to trimethylsilated (TMS) forms. The samples were then aliquoted into an autosampler tray for GC-MS analysis. An Agilent GC 7890A coupled with a single quadrupole MSD 5975C (Agilent Technologies) was used for collection of GC-MS data. Data was collected over a mass range of 50-550 m/z. A standard mixture of fatty acid methyl esters (FAMEs) (C8-C28) was analyzed with samples for RI alignment. The GC oven was held at 60ºC for 1 min after injection followed by a temperature increase by 10 ºC min-1 to a maximum of 325 ºC at which point it will be held for 5 min. All raw data can be found on the MassIVE archive with ID MSV000089933.