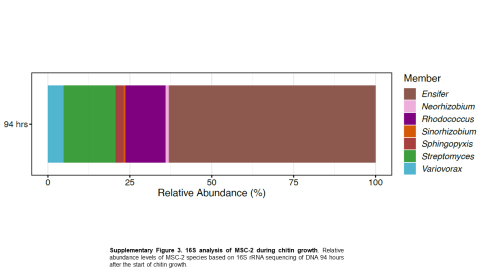
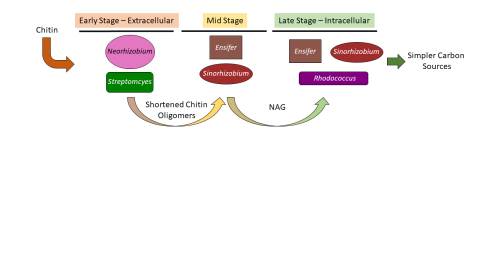
Please cite as : McClure R.S., Y. Farris, R.E. Danczak, W.C. Nelson, H. Song, A. Kessler, and J. Lee, et al. 2022. 16s data from MSC-2 growth. [Data Set] PNNL DataHub. https://data.pnnl.gov/group/nodes/dataset/33231 16s data from MSC-2 growth 3 fastq of 16s amplicon data of MSC2 1 csv file of raw...
Filter results
Content type
Tags
- (-) Chitin (2)
- Omics (12)
- PerCon SFA (9)
- High Throughput Sequencing (8)
- Genomics (7)
- Machine Learning (7)
- Type 1 Diabetes (6)
- Autoimmunity (5)
- Mass Spectrometry (5)
- Sequencer System (5)
- Synthetic (5)
- Synthetic Biology (5)
- Biomarkers (4)
- Molecular Profiling (4)
- Predictive Modeling (4)
- RNA Sequence Analysis (4)
- Mass spectrometry-based Omics (3)
- Software Data Analysis (3)
- Statistical Expression Analysis (3)
- Amplicon Sequencing (2)
- Biological and Environmental Research (2)
- Imaging (2)
- Long Read Sequencer (2)
- Mass Spectrometer (2)
- Mass spectrometry data (2)
- Proteomics (2)
- Soil (2)
- Soil Microbiology (2)
- Spectroscopy (2)
- Whole Genome Sequencing (2)
Showing 1 - 3 of 3
Dataset
Please cite as : McClure R.S., Y. Farris, R.E. Danczak, W.C. Nelson, H. Song, A. Kessler, and J. Lee, et al. 2022. Metatranscriptomic data from MSC-2. [Data Set] PNNL DataHub. https://data.pnnl.gov/group/nodes/dataset/33232 Metatranscriptomic data from MSC-2 12 fastq files (6 forward read, 6 reverse...
Category
pmartR Software Overview The pmartR package provides a single software tool for QC (filtering and normalization), exploratory data analysis (EDA), and statistical analysis (robust to missing data) and includes numerous visualization capabilities of mass spectrometry (MS) omics data (proteomic...