pmartR Software Overview The pmartR package provides a single software tool for QC (filtering and normalization), exploratory data analysis (EDA), and statistical analysis (robust to missing data) and includes numerous visualization capabilities of mass spectrometry (MS) omics data (proteomic...
Filter results
Category
- (-) Ecosystem Science (2)
- (-) Computational Research (1)
- (-) Materials Science (1)
- Scientific Discovery (42)
- Biology (40)
- Earth System Science (37)
- Microbiome Science (11)
- Integrative Omics (9)
- Human Health (6)
- Chemical & Biological Signatures Science (1)
- Chemistry (1)
- National Security (1)
- Plant Science (1)
- Weapons of Mass Effect (1)
Content type
Tags
- (-) Omics (3)
- Type 1 Diabetes (6)
- Autoimmunity (5)
- Machine Learning (5)
- Biomarkers (4)
- Molecular Profiling (4)
- Mass spectrometry-based Omics (3)
- Predictive Modeling (3)
- Software Data Analysis (3)
- Alternative Splicing (2)
- DNA Sequence Analysis (2)
- Functional Annotation Analysis (2)
- Kmers (2)
- Mass Spectrometry (2)
- Polymer Materials (2)
- Python (2)
- RNA Sequence Analysis (2)
- Snakemake (2)
- High-Performance Computing (1)
- Imaging (1)
- Mass Spectrometer (1)
- Microbeam (1)
- Microscopy (1)
- Output Databases (1)
- Spectroscopy (1)
- Statistics (1)
- ToF-SIMS (1)
- underwater video (1)
- X-Ray Diffraction (1)
- XANES (1)
The IONTOF TOF.SIMS 5 data source is a time-of-flight secondary ion mass spectrometer and powerful surface analysis tool used to investigate scientific questions in biological, environmental, and energy research. Among the most sensitive of surface analysis tools, it uses a high-vacuum technique...
Stanford Synchrotron Radiation Lightsource Experimental Station 14-3b is a bending magnet side station dedicated to X-Ray Imaging and Micro X-Ray Absorption Spectroscopy of biological, biomedical, materials, and geological samples. Station 14-3b is equipped with specialized instrumentation for XRF...
Category
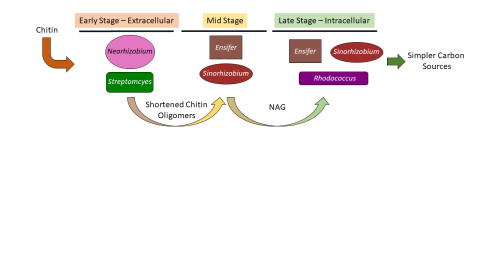
Please cite as : McClure R.S., Y. Farris, R.E. Danczak, W.C. Nelson, H. Song, A. Kessler, and J. Lee, et al. 2022. Metatranscriptomic data from MSC-2. [Data Set] PNNL DataHub. https://data.pnnl.gov/group/nodes/dataset/33232 Metatranscriptomic data from MSC-2 12 fastq files (6 forward read, 6 reverse...