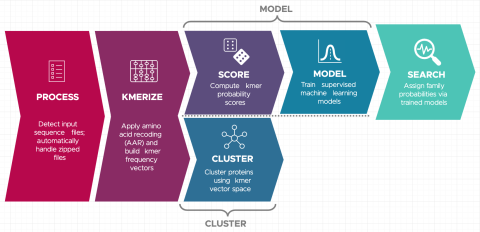
Last updated on 2023-02-23T19:37:46+00:00 by LN Anderson Snekmer: A scalable pipeline for protein sequence fingerprinting using amino acid recoding (AAR) Snekmer is a software package designed to reduce the representation of protein sequences by combining amino acid reduction (AAR) with the kmer...
Filter results
Category
Content type
Dataset Type
Tags
- (-) Machine Learning (2)
- (-) Kmers (1)
- Type 1 Diabetes (2)
- Alternative Splicing (1)
- Autoimmunity (1)
- DNA Sequence Analysis (1)
- EBC (1)
- Functional Annotation Analysis (1)
- High-Performance Computing (1)
- Mass Spectrometry (1)
- Omics (1)
- Predictive Modeling (1)
- Proteomics (1)
- Python (1)
- Quantification (1)
- RNA Sequence Analysis (1)
- Snakemake (1)
- Software Data Analysis (1)
- Statistics (1)
- TMT (1)
Showing 1 - 3 of 3
Software
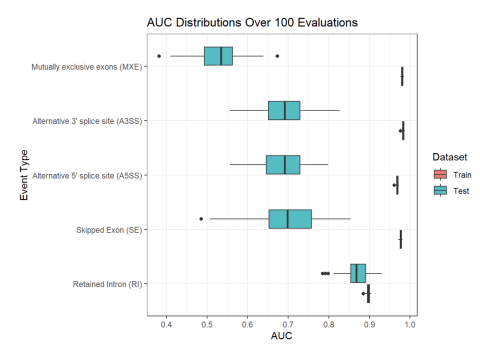
Inclusion levels of alternative splicing (AS) events of five different varieties (i.e. skipped exon (SE), retained intron (RI), alternative 5’ splice site (A5SS), alternative 3’ splice site (A3SS), and mutually exclusive exons (MXE)) were measured in human blood samples from two separate cohorts of...
Comprised of 6,426 sample runs, The Environmental Determinants of Diabetes in the Young (TEDDY) proteomics validation study constitutes one of the largest targeted proteomics studies in the literature to date. Making quality control (QC) and donor sample data available to researchers aligns with...