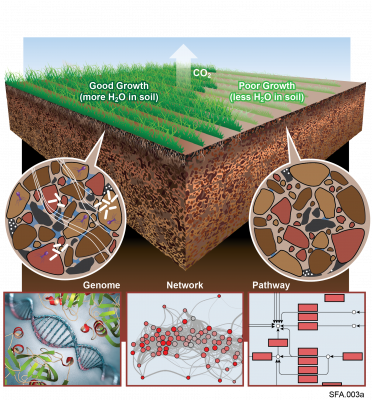
"Visualizing the Hidden Half: Plant-Microbe Interactions in the Rhizosphere" Plant roots and the associated rhizosphere constitute a dynamic environment that fosters numerous intra- and interkingdom interactions, including metabolite exchange between plants and soil mediated by root exudates and the...
Filter results
Content type
Tags
- (-) Chitin (2)
- (-) Spectroscopy (2)
- Omics (11)
- PerCon SFA (9)
- High Throughput Sequencing (8)
- Genomics (7)
- Machine Learning (6)
- Type 1 Diabetes (6)
- Autoimmunity (5)
- Mass Spectrometry (5)
- Sequencer System (5)
- Synthetic Biology (5)
- Biomarkers (4)
- Molecular Profiling (4)
- RNA Sequence Analysis (4)
- Mass spectrometry-based Omics (3)
- Predictive Modeling (3)
- Software Data Analysis (3)
- Statistical Expression Analysis (3)
- Amplicon Sequencing (2)
- Biological and Environmental Research (2)
- DNA Sequence Analysis (2)
- Functional Annotation Analysis (2)
- Imaging (2)
- Long Read Sequencer (2)
- Mass Spectrometer (2)
- Mass spectrometry data (2)
- Proteomics (2)
- Soil (2)
- Whole Genome Sequencing (2)
The Phenotypic Response of the Soil Microbiome to Environmental Perturbations Project (Soil Microbiome SFA) at Pacific Northwest National Laboratory is a Genomic Sciences Program Science Focus Area (SFA) Project operating under the Environmental Microbiome Science Research Area. The Soil Microbiome...
Datasets
23
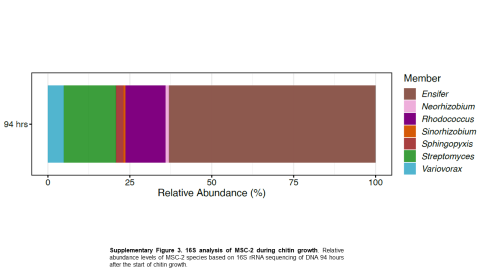
Please cite as : McClure R.S., Y. Farris, R.E. Danczak, W.C. Nelson, H. Song, A. Kessler, and J. Lee, et al. 2022. 16s data from MSC-2 growth. [Data Set] PNNL DataHub. https://data.pnnl.gov/group/nodes/dataset/33231 16s data from MSC-2 growth 3 fastq of 16s amplicon data of MSC2 1 csv file of raw...
Category
Please cite as : McClure R.S., Y. Farris, R.E. Danczak, W.C. Nelson, H. Song, A. Kessler, and J. Lee, et al. 2022. Metatranscriptomic data from MSC-2. [Data Set] PNNL DataHub. https://data.pnnl.gov/group/nodes/dataset/33232 Metatranscriptomic data from MSC-2 12 fastq files (6 forward read, 6 reverse...