A major challenge in biotechnology and biomanufacturing is the identification of a set of biomarkers for perturbations and metabolites of interest. Here, we develop a data-driven, transcriptome-wide approach to rank perturbation-inducible genes from time-series RNA sequencing data for the discovery...
Filter results
Category
- (-) Microbiome Science (2)
- (-) Computational Mathematics & Statistics (1)
- (-) Data Analytics & Machine Learning (1)
- (-) Earth System Science (1)
- Scientific Discovery (8)
- Biology (7)
- Computational Research (5)
- Data Analytics & Machine Learning (5)
- Human Health (5)
- Computational Mathematics & Statistics (1)
- Computing & Analytics (1)
- Ecosystem Science (1)
- National Security (1)
Content type
Tags
- (-) Machine Learning (3)
- Omics (22)
- Soil Microbiology (21)
- sequencing (13)
- Genomics (12)
- High Throughput Sequencing (9)
- Metagenomics (9)
- PerCon SFA (9)
- Microbiome (8)
- Fungi (6)
- Imaging (6)
- Mass Spectrometry (6)
- Mass Spectrometer (5)
- Sequencer System (5)
- Synthetic Biology (5)
- metagenomics (4)
- Microscopy (4)
- Proteomics (4)
- Sequencing (4)
- soil microbiology (4)
- Spectroscopy (4)
- Amplicon Sequencing (3)
- Climate Change (3)
- IAREC (3)
- Mass spectrometry data (3)
- metabolomics (3)
- Viruses (3)
- Whole Genome Sequencing (3)
- microbiome stability (2)
- species volatility (2)
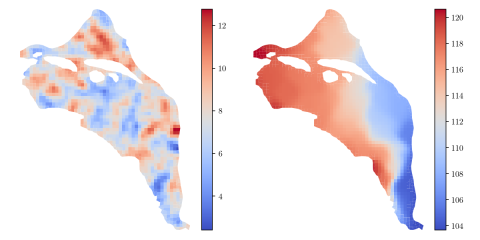
HDF5 file containing 10,000 hydraulic transmissivity inputs and the corresponding hydraulic pressure field outputs for a two-dimensional saturated flow model of the Hanford Site. The inputs are generated by sampling a 1,000-dimensional Kosambi-Karhunen-Loève (KKL) model of the transmissivity field...
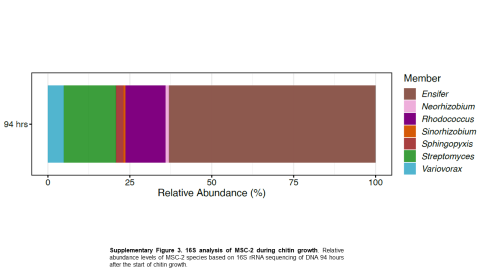
Please cite as : McClure R.S., Y. Farris, R.E. Danczak, W.C. Nelson, H. Song, A. Kessler, and J. Lee, et al. 2022. 16s data from MSC-2 growth. [Data Set] PNNL DataHub. https://data.pnnl.gov/group/nodes/dataset/33231 16s data from MSC-2 growth 3 fastq of 16s amplicon data of MSC2 1 csv file of raw...
Category
Please cite as : McClure R.S., Y. Farris, R.E. Danczak, W.C. Nelson, H. Song, A. Kessler, and J. Lee, et al. 2022. Metatranscriptomic data from MSC-2. [Data Set] PNNL DataHub. https://data.pnnl.gov/group/nodes/dataset/33232 Metatranscriptomic data from MSC-2 12 fastq files (6 forward read, 6 reverse...
Category
Comprised of 6,426 sample runs, The Environmental Determinants of Diabetes in the Young (TEDDY) proteomics validation study constitutes one of the largest targeted proteomics studies in the literature to date. Making quality control (QC) and donor sample data available to researchers aligns with...