Christine H Chang, William C Nelson, Abby Jerger, Aaron T Wright, Robert G Egbert, Jason E McDermott, Snekmer: a scalable pipeline for protein sequence fingerprinting based on amino acid recoding, Bioinformatics Advances , Volume 3, Issue 1, 2023, vbad005, https://doi.org/10.1093/bioadv/vbad005...
Filter results
Content type
Tags
- (-) Alternative Splicing (2)
- (-) Functional Annotation Analysis (2)
- Machine Learning (6)
- Type 1 Diabetes (6)
- Autoimmunity (5)
- Biomarkers (4)
- Molecular Profiling (4)
- Predictive Modeling (4)
- Mass spectrometry-based Omics (3)
- Omics (3)
- Software Data Analysis (3)
- DNA Sequence Analysis (2)
- Kmers (2)
- Mass Spectrometry (2)
- Proteomics (2)
- Python (2)
- RNA Sequence Analysis (2)
- Snakemake (2)
- EBC (1)
- Exhaled Breath Condensate (1)
- High-Performance Computing (1)
- Hydropower (1)
- Imaging (1)
- Marine and Hydrokinetic (1)
- Output Databases (1)
- Quantification (1)
- Spectroscopy (1)
- Statistics (1)
- TMT (1)
- underwater video (1)
The Human Islet Research Network (HIRN) is a large consortia with many research projects focused on understanding how beta cells are lost in type 1 diabetics (T1D) with a goal of finding how to protect against or replace the loss of functional beta cells. The consortia has multiple branches of...
Datasets
0
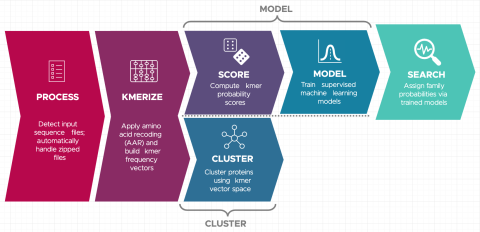
Last updated on 2023-02-23T19:37:46+00:00 by LN Anderson Snekmer: A scalable pipeline for protein sequence fingerprinting using amino acid recoding (AAR) Snekmer is a software package designed to reduce the representation of protein sequences by combining amino acid reduction (AAR) with the kmer...
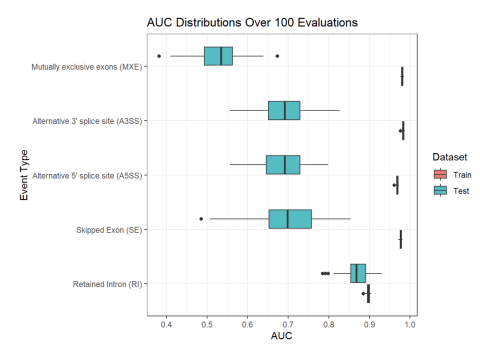
Inclusion levels of alternative splicing (AS) events of five different varieties (i.e. skipped exon (SE), retained intron (RI), alternative 5’ splice site (A5SS), alternative 3’ splice site (A3SS), and mutually exclusive exons (MXE)) were measured in human blood samples from two separate cohorts of...