Omics-LHV, West Nile Experiment WGCN004 The purpose of this West Nile experiment was to obtain samples for omics analysis in primary mouse granule neuron cells infected with wild type West Nile virus (WNV-NY99 clone 382, WNVWT) and mutant virus (WNVE218A). Overall Design: Granule cell neurons from...
Filter results
Category
- (-) Integrative Omics (1)
- (-) National Security (1)
- Biology (30)
- Scientific Discovery (30)
- Human Health (28)
- Computational Research (5)
- Data Analytics & Machine Learning (5)
- Chemistry (1)
- Computational Mathematics & Statistics (1)
- Computational Mathematics & Statistics (1)
- Computing & Analytics (1)
- Data Analytics & Machine Learning (1)
- Microbiome Science (1)
Content type
Tags
- Virology (57)
- Immune Response (53)
- Time Sampled Measurement Datasets (50)
- Gene expression profile data (47)
- Differential Expression Analysis (46)
- Homo sapiens (34)
- Mass spectrometry data (30)
- Multi-Omics (30)
- MERS-CoV (18)
- Mus musculus (18)
- West Nile virus (13)
- Ebola (11)
- Influenza A (11)
- Omics (9)
- Resource Metadata (9)
- Mass Spectrometry (7)
- Human Interferon (6)
- Microarray (6)
- Omics-LHV Project (6)
- Proteomics (6)
- Synthetic (5)
- Mass Spectrometer (4)
- Imaging (3)
- metabolomics (3)
- Microscopy (3)
- Response to type I interferon (3)
- Cybersecurity (2)
- Electrical energy (2)
- Epigenetics (2)
- Lipidomics (2)
Showing 1 - 2 of 2
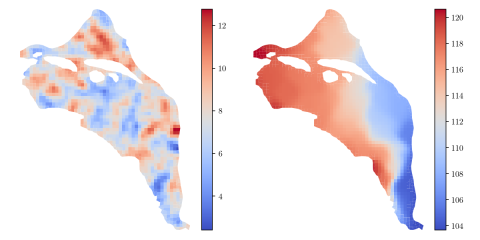
HDF5 file containing 10,000 hydraulic transmissivity inputs and the corresponding hydraulic pressure field outputs for a two-dimensional saturated flow model of the Hanford Site. The inputs are generated by sampling a 1,000-dimensional Kosambi-Karhunen-Loève (KKL) model of the transmissivity field...