Christine H Chang, William C Nelson, Abby Jerger, Aaron T Wright, Robert G Egbert, Jason E McDermott, Snekmer: a scalable pipeline for protein sequence fingerprinting based on amino acid recoding, Bioinformatics Advances , Volume 3, Issue 1, 2023, vbad005, https://doi.org/10.1093/bioadv/vbad005...
Filter results
Content type
Tags
- (-) Synthetic (5)
- (-) Kmers (2)
- Machine Learning (6)
- Type 1 Diabetes (6)
- Autoimmunity (5)
- Biomarkers (4)
- Molecular Profiling (4)
- Predictive Modeling (4)
- Mass spectrometry-based Omics (3)
- Software Data Analysis (3)
- Alternative Splicing (2)
- Cybersecurity (2)
- DNA Sequence Analysis (2)
- Electrical energy (2)
- Functional Annotation Analysis (2)
- Omics (2)
- Proteomics (2)
- Python (2)
- RNA Sequence Analysis (2)
- Snakemake (2)
- EBC (1)
- Exhaled Breath Condensate (1)
- High-Performance Computing (1)
- Marine and Hydrokinetic (1)
- Mass Spectrometry (1)
- Output Databases (1)
- Quantification (1)
- Statistics (1)
- TMT (1)
- underwater video (1)
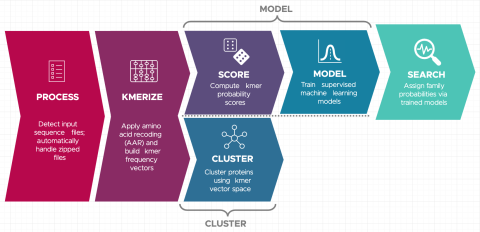
Last updated on 2023-02-23T19:37:46+00:00 by LN Anderson Snekmer: A scalable pipeline for protein sequence fingerprinting using amino acid recoding (AAR) Snekmer is a software package designed to reduce the representation of protein sequences by combining amino acid reduction (AAR) with the kmer...
This data is a model of synthetic adversarial activity surrounded by noise and was funded by DARPA. The various versions include gradually more complex networks of activities.
Category
Datasets
1
Category
Datasets
7
This data is a model of synthetic adversarial activity surrounded by noise and was funded by DARPA. The various versions include gradually more complex networks of activities.
Category
Datasets
1
Category
Datasets
1
This data was generated by the organization IvySys. Activities can be phone calls, transactions, or any other type of communications. Most of the files are of the type .edges, .rdf, or .csv; but all can be opened in a text editor. A good introduction to this data can be found in \Tutorial1\MAA...