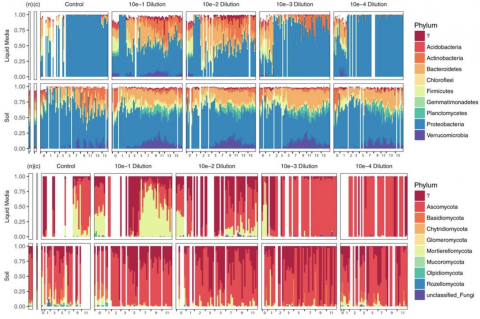
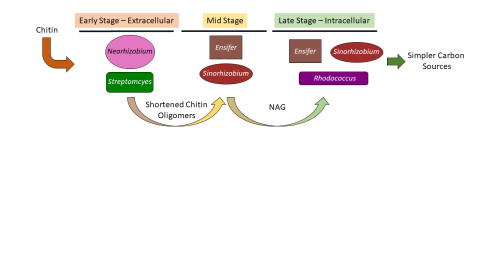
Please cite as : Zegeye E., C.J. Brislawn, Y. Farris, S.J. Fansler, K.S. Hofmockel, J.K. Jansson, and A.T. Wright, et al. 2019. WA-IsoC_NAG.1.0 (Amplicon 16S/ITS, WA). [Data Set] PNNL DataHub. https://dx.doi.org/10.25584/data.2019-02.700/1506698 Investigation of the successional dynamics of a soil...
Filter results
Content type
Tags
- (-) microbiome stability (1)
- (-) MultiSector Dynamics (1)
- (-) Soil (1)
- Soil Microbiology (13)
- sequencing (9)
- Metagenomics (7)
- Microbiome (4)
- Omics (4)
- Fungi (3)
- Genomics (3)
- IAREC (3)
- metagenomics (3)
- Sequencing (3)
- soil microbiology (3)
- Climate Change (2)
- Mass Spectrometry (2)
- omics (2)
- Amplicon Sequencing (1)
- Chitin (1)
- High Throughput Sequencing (1)
- Metatranscriptomic (1)
- Microscopy (1)
- mineral weathering (1)
- MPLEX (1)
- Output Databases (1)
- Software Data Analysis (1)
- species volatility (1)
- Spectroscopy (1)
- Viruses (1)
- Whole Genome Sequencing (1)
Showing 1 - 3 of 3
These GCAM v4.3 SSP-RCP-GCM Output Databases are made available under the Open Data Commons Attribution License: http://opendatacommons.org/licenses/by/1.0/ . GCAM v4.3 SSP-RCP-GCM plausible solution databases. Supplemental dataset to: Graham N.T., M.I. Hejazi, M. Chen, E. Davies, J.A. Edmonds, S.H...
Category
Please cite as : McClure R.S., Y. Farris, R.E. Danczak, W.C. Nelson, H. Song, A. Kessler, and J. Lee, et al. 2022. Metatranscriptomic data from MSC-2. [Data Set] PNNL DataHub. https://data.pnnl.gov/group/nodes/dataset/33232 Metatranscriptomic data from MSC-2 12 fastq files (6 forward read, 6 reverse...