"Visualizing the Hidden Half: Plant-Microbe Interactions in the Rhizosphere" Plant roots and the associated rhizosphere constitute a dynamic environment that fosters numerous intra- and interkingdom interactions, including metabolite exchange between plants and soil mediated by root exudates and the...
Filter results
Content type
Tags
- (-) Python (2)
- (-) Spectroscopy (1)
- Type 1 Diabetes (6)
- Autoimmunity (5)
- Machine Learning (5)
- Mass Spectrometry (5)
- Biomarkers (4)
- Molecular Profiling (4)
- Mass spectrometry-based Omics (3)
- Omics (3)
- Predictive Modeling (3)
- Software Data Analysis (3)
- Alternative Splicing (2)
- DNA Sequence Analysis (2)
- Functional Annotation Analysis (2)
- Kmers (2)
- PerCon SFA (2)
- RNA Sequence Analysis (2)
- Snakemake (2)
- High-Performance Computing (1)
- Hydropower (1)
- Imaging (1)
- Marine and Hydrokinetic (1)
- Mass Spectrometer (1)
- metabolomics (1)
- Output Databases (1)
- Proteomics (1)
- Statistics (1)
- ToF-SIMS (1)
- underwater video (1)
Showing 1 - 3 of 3
Publication
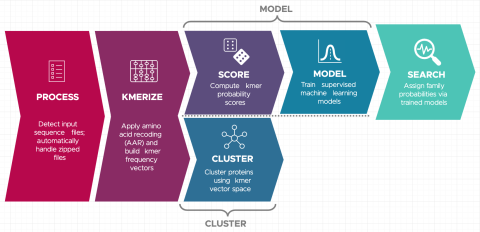
Christine H Chang, William C Nelson, Abby Jerger, Aaron T Wright, Robert G Egbert, Jason E McDermott, Snekmer: a scalable pipeline for protein sequence fingerprinting based on amino acid recoding, Bioinformatics Advances , Volume 3, Issue 1, 2023, vbad005, https://doi.org/10.1093/bioadv/vbad005...
Software
Last updated on 2023-02-23T19:37:46+00:00 by LN Anderson Snekmer: A scalable pipeline for protein sequence fingerprinting using amino acid recoding (AAR) Snekmer is a software package designed to reduce the representation of protein sequences by combining amino acid reduction (AAR) with the kmer...