MERS-CoV Experiment MDC001 Processed Omics Data Unavailable This experiment evaluated primary human dendritic cells infected with a wild type MERS-CoV (icMERS) virus. Related Experimental Data BioProject: PRJNA315103 GEO: GSE79172 (mRNA transcriptome response) Acknowledgment of Federal Funding The...
Filter results
Category
- Biology (3)
- Human Health (3)
- Scientific Discovery (3)
- Computational Research (2)
- Data Analytics & Machine Learning (2)
- National Security (2)
- Computational Mathematics & Statistics (1)
- Computational Mathematics & Statistics (1)
- Computing & Analytics (1)
- Cybersecurity (1)
- Data Analytics & Machine Learning (1)
- Distribution (1)
- Electric Grid Modernization (1)
- Energy Resiliency (1)
- Grid Analytics (1)
- Grid Cybersecurity (1)
- Integrative Omics (1)
Content type
Tags
- (-) Machine Learning (3)
- Virology (61)
- Immune Response (46)
- Time Sampled Measurement Datasets (43)
- Differential Expression Analysis (41)
- Gene expression profile data (40)
- Homo sapiens (28)
- Mass spectrometry data (25)
- Multi-Omics (23)
- MERS-CoV (16)
- Mus musculus (15)
- Viruses (14)
- Health (13)
- Virus (13)
- West Nile virus (11)
- Soil Microbiology (10)
- Ebola (9)
- Influenza A (7)
- Metagenomics (7)
- sequencing (7)
- Resource Metadata (6)
- Microarray (5)
- Human Interferon (4)
- Omics (4)
- Genomics (3)
- metagenomics (3)
- Microbiome (3)
- PerCon SFA (3)
- Sequencing (3)
- soil microbiology (3)
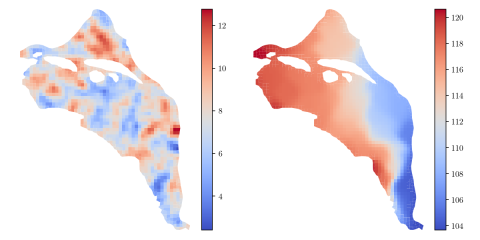
HDF5 file containing 10,000 hydraulic transmissivity inputs and the corresponding hydraulic pressure field outputs for a two-dimensional saturated flow model of the Hanford Site. The inputs are generated by sampling a 1,000-dimensional Kosambi-Karhunen-Loève (KKL) model of the transmissivity field...
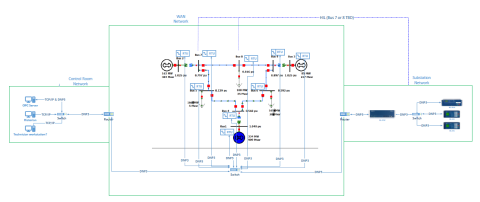
This dataset includes one baseline and three cybersecurity based scenarios utilizing the IEEE 9 Bus Model. This instantiation of the IEEE 9 model was built utilizing the OpalRT Simulator ePhasorsim module, with Bus 7 represented by hardware in the loop (HiL). The HiL was represented by two SEL351s...
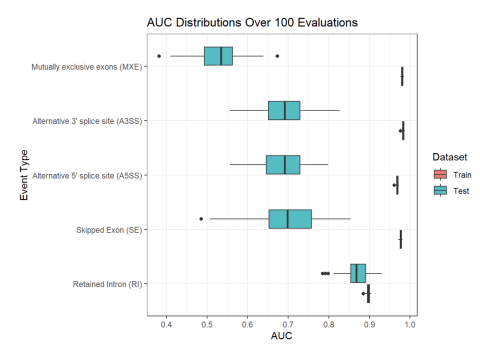
Inclusion levels of alternative splicing (AS) events of five different varieties (i.e. skipped exon (SE), retained intron (RI), alternative 5’ splice site (A5SS), alternative 3’ splice site (A3SS), and mutually exclusive exons (MXE)) were measured in human blood samples from two separate cohorts of...
Comprised of 6,426 sample runs, The Environmental Determinants of Diabetes in the Young (TEDDY) proteomics validation study constitutes one of the largest targeted proteomics studies in the literature to date. Making quality control (QC) and donor sample data available to researchers aligns with...