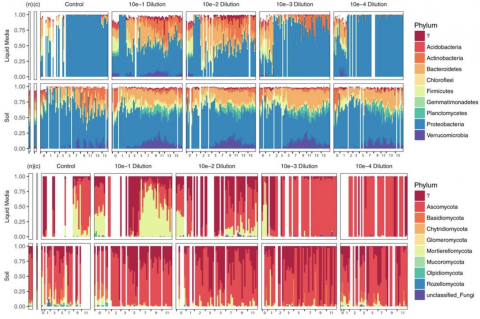
Please cite as : Zegeye E., C.J. Brislawn, Y. Farris, S.J. Fansler, K.S. Hofmockel, J.K. Jansson, and A.T. Wright, et al. 2019. WA-IsoC_NAG.1.0 (Amplicon 16S/ITS, WA). [Data Set] PNNL DataHub. https://dx.doi.org/10.25584/data.2019-02.700/1506698 Investigation of the successional dynamics of a soil...
Filter results
Content type
Tags
- (-) sequencing (2)
- Virology (50)
- Gene expression profile data (40)
- Immune Response (40)
- Time Sampled Measurement Datasets (37)
- Differential Expression Analysis (35)
- Homo sapiens (24)
- Mass spectrometry data (18)
- Multi-Omics (17)
- MERS-CoV (14)
- Mus musculus (13)
- West Nile virus (9)
- Health (8)
- Virus (8)
- Viruses (8)
- Ebola (7)
- Influenza A (7)
- Resource Metadata (6)
- Microarray (5)
- Human Interferon (4)
- Response to type I interferon (3)
- Soil Microbiology (3)
- Amplicon Sequencing (2)
- Response to type I & type II interferon (2)
- Viral Experiment (2)
- IAREC (1)
- Mass Spectrometry (1)
- Microbiome (1)
- microbiome stability (1)
- species volatility (1)
Showing 1 - 2 of 2
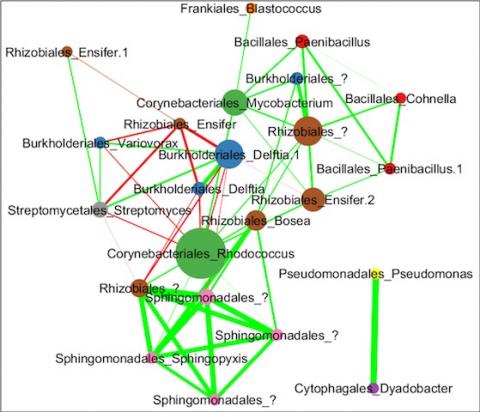
Please cite as : Anderson L.N., J.E. McDermott, and R.S. McClure. 2020. WA-IsoC_MSC1.1.0 (Amplicon 16S rRNA, WA). [Data Set] PNNL DataHub. https://doi.org/10.25584/WAIsoCMSC1/1635272 The soil microbiome is central to the cycling of carbon and other nutrients and to the promotion of plant growth...