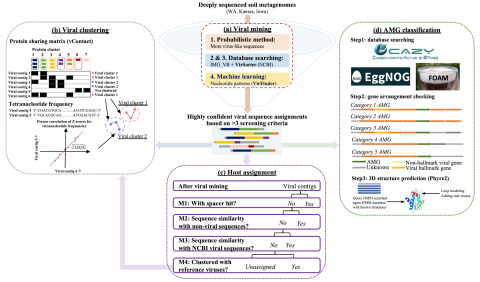
Viruses are abundant in soil, but the influence of the soil environment and climate on soil viruses remains unknown. Here we addressed this gap by comparing the diversity, abundance, lifestyle and metabolic potential of DNA viruses in three grassland soils with historical differences in average annual precipitation: Eastern Washington (WA), Kansas (KS), and Iowa (IA) . Deep metagenomes (>1 terabase each) were screened for viruses, resulting in 2,631 viral contigs, including 14 complete viral genomes. Identified soil viruses were primarily bacteriophages targeting dominant bacterial taxa and observed phage diversity correlated with host diversity. Viral community diversity and abundance were significantly higher in the arid WA location, compared to IA and KS. More lysogenic markers and fewer CRISPR spacer hits were found in WA, reflecting more lysogeny in the historically dry soil. More auxiliary metabolic genes (AMGs) were also detected in WA compared to the historically wetter locations. Across all sites the AMGs represented 18 pathways, suggesting viral contributions to carbon metabolism and energy acquisition in soil. Together these findings provide new knowledge of viruses in grassland soils and have value for predicting viral responses to changes in their environment.
Terabase soil metagenome samples were collected from Washington State University’s Irrigated Agriculture Research and Extension Center (IAREC), Konza Prairie Biological Station (KPBS), and Iowa State University’s Comparison of Biofuel Systems (COBS) and were collected from the same independent soil site collection tubes as the first Terabase Metagenome release (WA-TmG.1.0, KS-TmG.1.0, and IA-TmG.1.0).The versions described in this paper are the second version (2.0). Dataset downloads contain raw read sequencing files, assembly files, functional annotations, MIMS.me.soil.5.0 metadata information, and a dataset “Read Me” file.
Related Experimental Data
TmG.1.0 Metagenomes: 10.25584/WATmG1/1635002 | 10.25584/KSTmG1/1635004 | 10.25584/IATmG1/1635005
TmG.2.0 Metagenomes: 10.25584/WATmG2/1770324 | 10.25584/KSTmG2/1770332 | 10.25584/IATmG2/1770333