Citation: Gosline, S.J.C., Kim, D.N., Pande, P. et al. The Superfund Research Program Analytics Portal: linking environmental chemical exposure to biological phenotypes. Sci Data 10 , 151 (2023). https://doi.org/10.1038/s41597-023-02021-5 Funding Acknowledgments The research reported herein was...
Filter results
Content type
Tags
- (-) Kmers (2)
- (-) Toxicology (1)
- Health (8)
- Virology (8)
- Virus (8)
- Viruses (8)
- Mass Spectrometry (3)
- Software Data Analysis (3)
- DNA Sequence Analysis (2)
- Functional Annotation Analysis (2)
- Python (2)
- RNA Sequence Analysis (2)
- Snakemake (2)
- Data Analytics (1)
- Fish Detection (1)
- Host Immune Response (1)
- Hydropower (1)
- Lethal Human Viruses (1)
- Lipidomics (1)
- Marine and Hydrokinetic (1)
- Microarray (1)
- Multi-Omics Viral Data (1)
- Omics (1)
- Polycyclic Aromatic Hydrocarbons (1)
- Processed Data (1)
- Statistics (1)
- Superfund Hazardous Substances (1)
- underwater video (1)
Showing 1 - 3 of 3
Publication
Christine H Chang, William C Nelson, Abby Jerger, Aaron T Wright, Robert G Egbert, Jason E McDermott, Snekmer: a scalable pipeline for protein sequence fingerprinting based on amino acid recoding, Bioinformatics Advances , Volume 3, Issue 1, 2023, vbad005, https://doi.org/10.1093/bioadv/vbad005...
Software
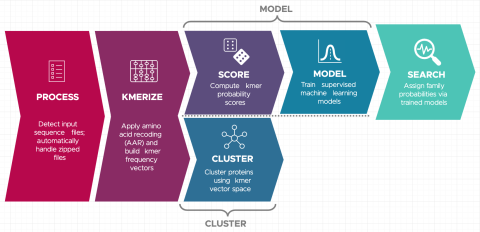
Last updated on 2023-02-23T19:37:46+00:00 by LN Anderson Snekmer: A scalable pipeline for protein sequence fingerprinting using amino acid recoding (AAR) Snekmer is a software package designed to reduce the representation of protein sequences by combining amino acid reduction (AAR) with the kmer...