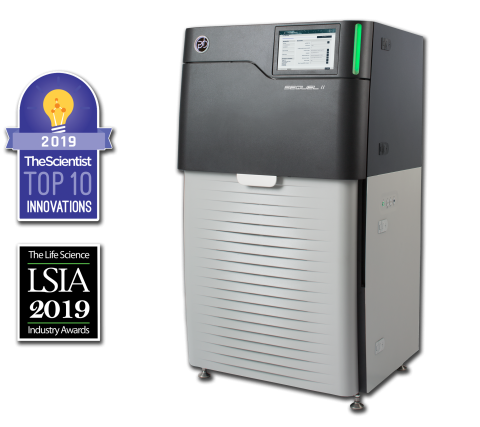
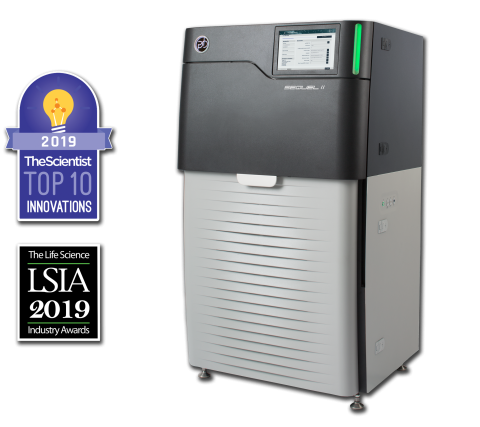
The Sequel II System Sequencer is a high-throughput DNA sequencer machine developed and manufactured by PacBio , and is designed for high throughput, production-scale sequencing laboratories. Originally released in 2015, the Sequel system provides Single Molecule, Real-Time (SMRT) sequencing core...
Filter results
Category
- (-) Human Health (5)
- Biology (20)
- Scientific Discovery (20)
- Earth System Science (17)
- Microbiome Science (5)
- Computational Research (2)
- Data Analytics & Machine Learning (2)
- Computational Mathematics & Statistics (1)
- Computational Mathematics & Statistics (1)
- Computing & Analytics (1)
- Data Analytics & Machine Learning (1)
- Integrative Omics (1)
- National Security (1)
Content type
Dataset Type
Tags
- (-) Sequencer System (3)
- (-) Machine Learning (2)
- Virology (63)
- Immune Response (48)
- Time Sampled Measurement Datasets (45)
- Gene expression profile data (42)
- Differential Expression Analysis (41)
- Homo sapiens (30)
- Mass spectrometry data (26)
- Multi-Omics (23)
- MERS-CoV (16)
- Mus musculus (15)
- Health (13)
- Virus (13)
- Viruses (13)
- West Nile virus (11)
- Ebola (9)
- Influenza A (9)
- Resource Metadata (8)
- Microarray (6)
- Omics (5)
- Human Interferon (4)
- Genomics (3)
- High Throughput Sequencing (3)
- Proteomics (3)
- Response to type I interferon (3)
- Epigenetics (2)
- Long Read Sequencer (2)
- Mass Spectrometer (2)
- Viral Experiment (2)
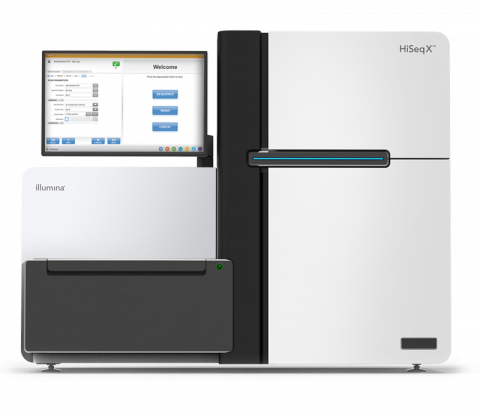
The Illumina HiSeq X System Sequencer is a high-throughput DNA sequencer machine developed and manufactured by Illumina , and is designed for high throughput, production-scale sequencing laboratories. Built off the HiSeq 2500 System, harnessing the patterned flow cell technology originally developed...
The Sequel II System Sequencer is a high-throughput DNA sequencer machine developed and manufactured by PacBio , and is designed for high throughput, production-scale sequencing laboratories. Originally released in 2015, the Sequel system provides Single Molecule, Real-Time (SMRT) sequencing core...
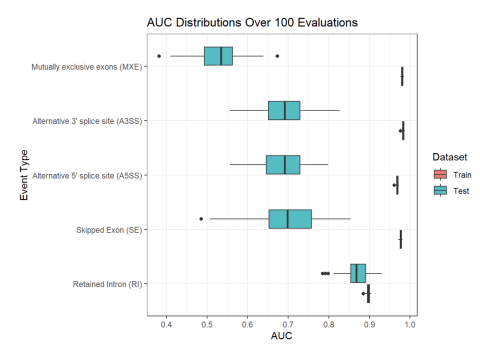
Inclusion levels of alternative splicing (AS) events of five different varieties (i.e. skipped exon (SE), retained intron (RI), alternative 5’ splice site (A5SS), alternative 3’ splice site (A3SS), and mutually exclusive exons (MXE)) were measured in human blood samples from two separate cohorts of...
Comprised of 6,426 sample runs, The Environmental Determinants of Diabetes in the Young (TEDDY) proteomics validation study constitutes one of the largest targeted proteomics studies in the literature to date. Making quality control (QC) and donor sample data available to researchers aligns with...