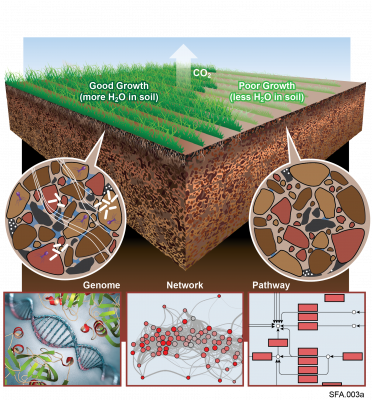
"Visualizing the Hidden Half: Plant-Microbe Interactions in the Rhizosphere" Plant roots and the associated rhizosphere constitute a dynamic environment that fosters numerous intra- and interkingdom interactions, including metabolite exchange between plants and soil mediated by root exudates and the...
Filter results
Content type
Tags
- (-) Functional Annotation Analysis (2)
- (-) Kmers (2)
- (-) Spectroscopy (2)
- Omics (11)
- PerCon SFA (9)
- High Throughput Sequencing (8)
- Genomics (7)
- Machine Learning (6)
- Type 1 Diabetes (6)
- Autoimmunity (5)
- Sequencer System (5)
- Synthetic Biology (5)
- Biomarkers (4)
- Mass Spectrometry (4)
- Molecular Profiling (4)
- RNA Sequence Analysis (4)
- Mass spectrometry-based Omics (3)
- Predictive Modeling (3)
- Software Data Analysis (3)
- Statistical Expression Analysis (3)
- Amplicon Sequencing (2)
- Biological and Environmental Research (2)
- DNA Sequence Analysis (2)
- Imaging (2)
- Long Read Sequencer (2)
- Mass spectrometry data (2)
- Proteomics (2)
- Python (2)
- Soil Microbiology (2)
- Whole Genome Sequencing (2)
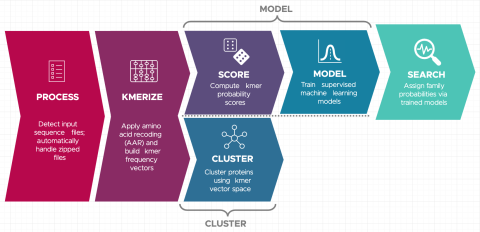
Christine H Chang, William C Nelson, Abby Jerger, Aaron T Wright, Robert G Egbert, Jason E McDermott, Snekmer: a scalable pipeline for protein sequence fingerprinting based on amino acid recoding, Bioinformatics Advances , Volume 3, Issue 1, 2023, vbad005, https://doi.org/10.1093/bioadv/vbad005...
Last updated on 2023-02-23T19:37:46+00:00 by LN Anderson Snekmer: A scalable pipeline for protein sequence fingerprinting using amino acid recoding (AAR) Snekmer is a software package designed to reduce the representation of protein sequences by combining amino acid reduction (AAR) with the kmer...
The Phenotypic Response of the Soil Microbiome to Environmental Perturbations Project (Soil Microbiome SFA) at Pacific Northwest National Laboratory is a Genomic Sciences Program Science Focus Area (SFA) Project operating under the Environmental Microbiome Science Research Area. The Soil Microbiome...
Datasets
23