Pending Review Microbiomes contribute to multiple ecosystem services by transforming organic matter in soil. Extreme shifts in the environment, such as drying-rewetting cycles during drought, can impact microbial metabolism of organic matter by altering their physiology and function. These...
Filter results
Content type
Tags
- (-) metabolomics (3)
- (-) Output Databases (2)
- Omics (22)
- Soil Microbiology (21)
- sequencing (13)
- Genomics (12)
- High Throughput Sequencing (9)
- Metagenomics (9)
- PerCon SFA (9)
- Microbiome (8)
- Fungi (6)
- Imaging (6)
- Mass Spectrometry (6)
- Mass Spectrometer (5)
- Sequencer System (5)
- Synthetic Biology (5)
- metagenomics (4)
- Microscopy (4)
- Proteomics (4)
- Sequencing (4)
- soil microbiology (4)
- Spectroscopy (4)
- Amplicon Sequencing (3)
- Climate Change (3)
- IAREC (3)
- Mass spectrometry data (3)
- Viruses (3)
- Whole Genome Sequencing (3)
- microbiome stability (2)
- species volatility (2)
Fusarium sp. DS682 Proteogenomics Statistical Data Analysis of SFA dataset download: 10.25584/KSOmicsFspDS682/1766303 . GitHub Repository Source: https://github.com/lmbramer/Fusarium-sp.-DS-682-Proteogenomics MaxQuant Export Files (txt) Trelliscope Boxplots (jsonp) Fusarium Report (.Rmd, html)...
The Thermo Scientific™ Q Exactive™ GC hybrid quadrupole Orbitrap Mass Spectrometer provides an unmatched combination of sensitivity, mass-accuracy and resolving power in a single analysis for the highest confidence in compound discovery, identification, and quantitation. With best-in-class...
Category
Bruker Daltonics SolariX Magnetic Resonance Mass Spectrometry (MRMS) instruments are available for different magnetic field strengths of 7T, 12T and 15T. The SolariX XR and 2xR instruments use magnetron control technology and a newly developed, high sensitivity, low noise preamplifier to exploit...
Category
These GCAM v4.3 SSP-RCP-GCM Output Databases are made available under the Open Data Commons Attribution License: http://opendatacommons.org/licenses/by/1.0/ . GCAM v4.3 SSP-RCP-GCM plausible solution databases. Supplemental dataset to: Graham N.T., M.I. Hejazi, M. Chen, E. Davies, J.A. Edmonds, S.H...
Category
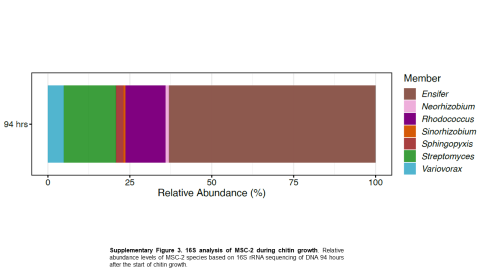
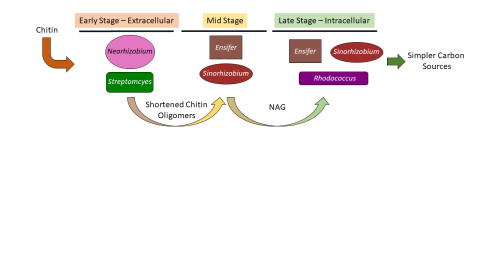
Please cite as : McClure R.S., Y. Farris, R.E. Danczak, W.C. Nelson, H. Song, A. Kessler, and J. Lee, et al. 2022. 16s data from MSC-2 growth. [Data Set] PNNL DataHub. https://data.pnnl.gov/group/nodes/dataset/33231 16s data from MSC-2 growth 3 fastq of 16s amplicon data of MSC2 1 csv file of raw...
Category
Please cite as : McClure R.S., Y. Farris, R.E. Danczak, W.C. Nelson, H. Song, A. Kessler, and J. Lee, et al. 2022. Metatranscriptomic data from MSC-2. [Data Set] PNNL DataHub. https://data.pnnl.gov/group/nodes/dataset/33232 Metatranscriptomic data from MSC-2 12 fastq files (6 forward read, 6 reverse...