A major challenge in biotechnology and biomanufacturing is the identification of a set of biomarkers for perturbations and metabolites of interest. Here, we develop a data-driven, transcriptome-wide approach to rank perturbation-inducible genes from time-series RNA sequencing data for the discovery...
Filter results
Category
- (-) Microbiome Science (3)
- (-) Ecosystem Science (2)
- Scientific Discovery (14)
- Biology (12)
- Human Health (7)
- Earth System Science (6)
- Computational Research (5)
- Data Analytics & Machine Learning (5)
- Integrative Omics (5)
- Chemistry (1)
- Computational Mathematics & Statistics (1)
- Computational Mathematics & Statistics (1)
- Computing & Analytics (1)
- Data Analytics & Machine Learning (1)
- National Security (1)
Content type
Tags
- (-) Mass Spectrometer (2)
- (-) Soil (2)
- (-) Machine Learning (1)
- Omics (10)
- PerCon SFA (9)
- High Throughput Sequencing (8)
- Genomics (7)
- Sequencer System (5)
- Synthetic Biology (5)
- Mass Spectrometry (4)
- A. pittii SO1 (2)
- Amplicon Sequencing (2)
- Biological and Environmental Research (2)
- Chitin (2)
- Imaging (2)
- Long Read Sequencer (2)
- Mass spectrometry data (2)
- Proteomics (2)
- RNA Sequence Analysis (2)
- Sorghum bicolor (2)
- Spectroscopy (2)
- Statistical Expression Analysis (2)
- Whole Genome Sequencing (2)
- Data Analysis (1)
- DOE (1)
- Lipidomics (1)
- metabolomics (1)
- Output Databases (1)
- Soil Microbiology (1)
- ToF-SIMS (1)
Last updated on 2024-03-03T02:26:52+00:00 by LN Anderson The Thermo Scientific™ Q Exactive™ Plus Mass Spectrometer benchtop LC-MS/MS system combines quadruple precursor ion selection with high-resolution, accurate-mass (HRAM) Orbitrap detection to deliver exceptional performance and versatility...
The IONTOF TOF.SIMS 5 data source is a time-of-flight secondary ion mass spectrometer and powerful surface analysis tool used to investigate scientific questions in biological, environmental, and energy research. Among the most sensitive of surface analysis tools, it uses a high-vacuum technique...
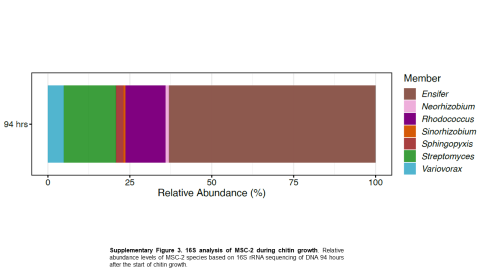
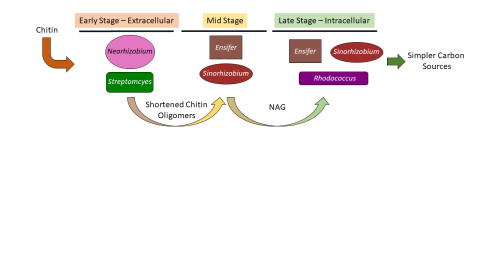
Please cite as : McClure R.S., Y. Farris, R.E. Danczak, W.C. Nelson, H. Song, A. Kessler, and J. Lee, et al. 2022. 16s data from MSC-2 growth. [Data Set] PNNL DataHub. https://data.pnnl.gov/group/nodes/dataset/33231 16s data from MSC-2 growth 3 fastq of 16s amplicon data of MSC2 1 csv file of raw...
Category
Please cite as : McClure R.S., Y. Farris, R.E. Danczak, W.C. Nelson, H. Song, A. Kessler, and J. Lee, et al. 2022. Metatranscriptomic data from MSC-2. [Data Set] PNNL DataHub. https://data.pnnl.gov/group/nodes/dataset/33232 Metatranscriptomic data from MSC-2 12 fastq files (6 forward read, 6 reverse...