"Moisture modulates soil reservoirs of active DNA and RNA viruses" Soil is known to harbor viruses, but the majority are uncharacterized and their responses to environmental changes are unknown. Here, we used a multi-omics approach (metagenomics, metatranscriptomics and metaproteomics) to detect...
Filter results
Content type
Tags
- (-) Data Analysis (1)
- (-) Metagenomics (1)
- Omics (10)
- PerCon SFA (9)
- High Throughput Sequencing (8)
- Genomics (7)
- Sequencer System (5)
- Synthetic Biology (5)
- Mass Spectrometry (3)
- A. pittii SO1 (2)
- Amplicon Sequencing (2)
- Biological and Environmental Research (2)
- Imaging (2)
- Long Read Sequencer (2)
- Mass spectrometry data (2)
- Proteomics (2)
- RNA Sequence Analysis (2)
- Soil Microbiology (2)
- Sorghum bicolor (2)
- Spectroscopy (2)
- Statistical Expression Analysis (2)
- Whole Genome Sequencing (2)
- DOE (1)
- Lipidomics (1)
- Machine Learning (1)
- Mass Spectrometer (1)
- metabolomics (1)
- Output Databases (1)
- Transcriptomics (1)
- Viruses (1)
Showing 1 - 3 of 3
Project
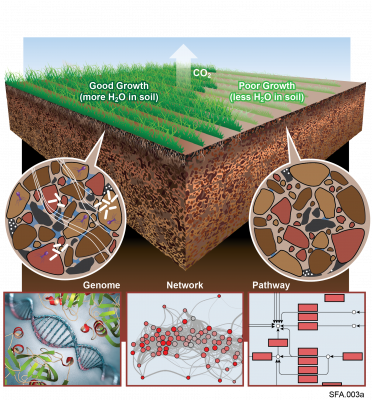
The Phenotypic Response of the Soil Microbiome to Environmental Perturbations Project (Soil Microbiome SFA) at Pacific Northwest National Laboratory is a Genomic Sciences Program Science Focus Area (SFA) Project operating under the Environmental Microbiome Science Research Area. The Soil Microbiome...
Datasets
23
Fusarium sp. DS682 Proteogenomics Statistical Data Analysis of SFA dataset download: 10.25584/KSOmicsFspDS682/1766303 . GitHub Repository Source: https://github.com/lmbramer/Fusarium-sp.-DS-682-Proteogenomics MaxQuant Export Files (txt) Trelliscope Boxplots (jsonp) Fusarium Report (.Rmd, html)...