"Visualizing the Hidden Half: Plant-Microbe Interactions in the Rhizosphere" Plant roots and the associated rhizosphere constitute a dynamic environment that fosters numerous intra- and interkingdom interactions, including metabolite exchange between plants and soil mediated by root exudates and the...
Filter results
Category
- (-) Computational Research (6)
- (-) Plant Science (1)
- Scientific Discovery (33)
- Biology (32)
- Earth System Science (22)
- Integrative Omics (10)
- Microbiome Science (10)
- Human Health (9)
- Data Analytics & Machine Learning (3)
- Chemistry (2)
- Chemical & Biological Signatures Science (1)
- Ecosystem Science (1)
- Materials Science (1)
- National Security (1)
- Weapons of Mass Effect (1)
Content type
Tags
- (-) Mass spectrometry-based Omics (3)
- (-) Omics (2)
- (-) Snakemake (2)
- Type 1 Diabetes (6)
- Autoimmunity (5)
- Machine Learning (5)
- Biomarkers (4)
- Molecular Profiling (4)
- Predictive Modeling (3)
- Software Data Analysis (3)
- Alternative Splicing (2)
- DNA Sequence Analysis (2)
- Functional Annotation Analysis (2)
- Kmers (2)
- Mass Spectrometry (2)
- Python (2)
- RNA Sequence Analysis (2)
- Bacterial Persistence (1)
- Bacterial Signaling (1)
- Data Analysis (1)
- Fish Detection (1)
- High-Performance Computing (1)
- Hydropower (1)
- Imaging (1)
- Marine and Hydrokinetic (1)
- Output Databases (1)
- Spectroscopy (1)
- Statistics (1)
- Tomography (1)
- underwater video (1)
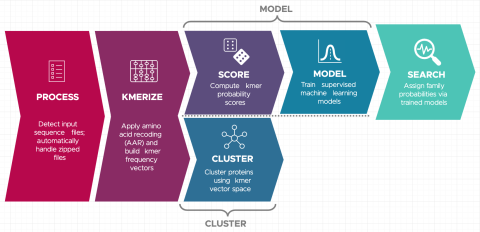
Christine H Chang, William C Nelson, Abby Jerger, Aaron T Wright, Robert G Egbert, Jason E McDermott, Snekmer: a scalable pipeline for protein sequence fingerprinting based on amino acid recoding, Bioinformatics Advances , Volume 3, Issue 1, 2023, vbad005, https://doi.org/10.1093/bioadv/vbad005...
pmartR Software Overview The pmartR package provides a single software tool for QC (filtering and normalization), exploratory data analysis (EDA), and statistical analysis (robust to missing data) and includes numerous visualization capabilities of mass spectrometry (MS) omics data (proteomic...
Last updated on 2023-02-23T19:37:46+00:00 by LN Anderson Snekmer: A scalable pipeline for protein sequence fingerprinting using amino acid recoding (AAR) Snekmer is a software package designed to reduce the representation of protein sequences by combining amino acid reduction (AAR) with the kmer...
The Environmental Determinants of Diabetes in the Young (TEDDY) study is searching for factors influencing the development of type 1 diabetes (T1D) in children. Research has shown that there are certain genes that correlate to higher risk of developing T1D, but not all children with these genes...
Datasets
1
The Diabetes Autoimmunity Study in the Young (DAISY) seeks to find environmental factors that can trigger the development of type 1 diabetes (T1D) in children. DAISY follows children with high-risk of developing T1D based on family history or genetic markers. Genes, diets, infections, and...
Datasets
1
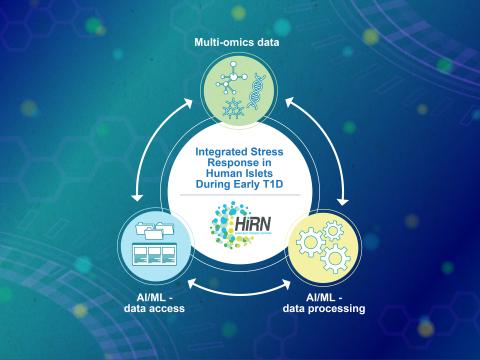
Machine learning is a core technology that is rapidly advancing within type 1 diabetes (T1D) research. Our Human Islet Research Network (HIRN) grant is studying early cellular response initiating β cell stress in T1D through the generation of heterogenous low- and high-throughput molecular...
Datasets
3