Christine H Chang, William C Nelson, Abby Jerger, Aaron T Wright, Robert G Egbert, Jason E McDermott, Snekmer: a scalable pipeline for protein sequence fingerprinting based on amino acid recoding, Bioinformatics Advances , Volume 3, Issue 1, 2023, vbad005, https://doi.org/10.1093/bioadv/vbad005...
Filter results
Content type
Tags
- (-) Kmers (2)
- (-) Cybersecurity (1)
- Type 1 Diabetes (6)
- Autoimmunity (5)
- Machine Learning (5)
- Biomarkers (4)
- Molecular Profiling (4)
- Mass spectrometry-based Omics (3)
- Predictive Modeling (3)
- Software Data Analysis (3)
- Alternative Splicing (2)
- DNA Sequence Analysis (2)
- Functional Annotation Analysis (2)
- Mass Spectrometry (2)
- Omics (2)
- Python (2)
- RNA Sequence Analysis (2)
- Snakemake (2)
- Chitin (1)
- Data Analysis (1)
- Electrical energy (1)
- Fish Detection (1)
- High-Performance Computing (1)
- Hydropower (1)
- Marine and Hydrokinetic (1)
- Mass Spectrometer (1)
- Output Databases (1)
- Statistics (1)
- ToF-SIMS (1)
- underwater video (1)
Showing 1 - 3 of 3
Publication
Software
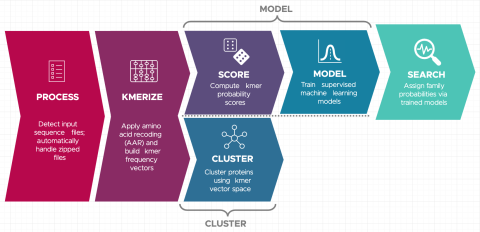
Last updated on 2023-02-23T19:37:46+00:00 by LN Anderson Snekmer: A scalable pipeline for protein sequence fingerprinting using amino acid recoding (AAR) Snekmer is a software package designed to reduce the representation of protein sequences by combining amino acid reduction (AAR) with the kmer...
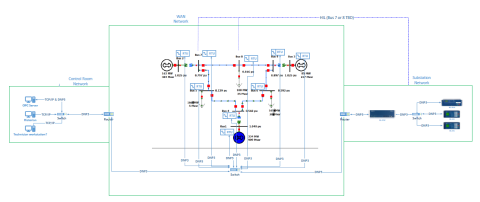
This dataset includes one baseline and three cybersecurity based scenarios utilizing the IEEE 9 Bus Model. This instantiation of the IEEE 9 model was built utilizing the OpalRT Simulator ePhasorsim module, with Bus 7 represented by hardware in the loop (HiL). The HiL was represented by two SEL351s...