Christine H Chang, William C Nelson, Abby Jerger, Aaron T Wright, Robert G Egbert, Jason E McDermott, Snekmer: a scalable pipeline for protein sequence fingerprinting based on amino acid recoding, Bioinformatics Advances , Volume 3, Issue 1, 2023, vbad005, https://doi.org/10.1093/bioadv/vbad005...
Filter results
Content type
Tags
- (-) Kmers (2)
- (-) RNA Sequence Analysis (2)
- (-) XANES (1)
- Type 1 Diabetes (6)
- Autoimmunity (5)
- Machine Learning (5)
- Biomarkers (4)
- Molecular Profiling (4)
- Mass spectrometry-based Omics (3)
- Predictive Modeling (3)
- Software Data Analysis (3)
- Alternative Splicing (2)
- DNA Sequence Analysis (2)
- Functional Annotation Analysis (2)
- Omics (2)
- Polymer Materials (2)
- Python (2)
- Snakemake (2)
- High-Performance Computing (1)
- Imaging (1)
- Marine and Hydrokinetic (1)
- Mass Spectrometry (1)
- Microbeam (1)
- Microscopy (1)
- Output Databases (1)
- Spectroscopy (1)
- Statistics (1)
- underwater video (1)
- X-Ray Diffraction (1)
- XRF (1)
Fusarium sp. DS682 Proteogenomics Statistical Data Analysis of SFA dataset download: 10.25584/KSOmicsFspDS682/1766303 . GitHub Repository Source: https://github.com/lmbramer/Fusarium-sp.-DS-682-Proteogenomics MaxQuant Export Files (txt) Trelliscope Boxplots (jsonp) Fusarium Report (.Rmd, html)...
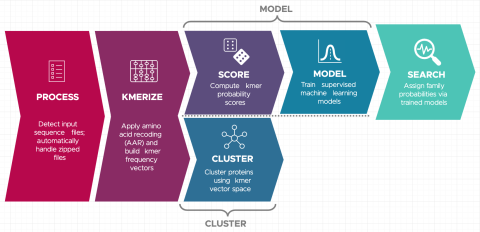
Last updated on 2023-02-23T19:37:46+00:00 by LN Anderson Snekmer: A scalable pipeline for protein sequence fingerprinting using amino acid recoding (AAR) Snekmer is a software package designed to reduce the representation of protein sequences by combining amino acid reduction (AAR) with the kmer...
Stanford Synchrotron Radiation Lightsource Experimental Station 14-3b is a bending magnet side station dedicated to X-Ray Imaging and Micro X-Ray Absorption Spectroscopy of biological, biomedical, materials, and geological samples. Station 14-3b is equipped with specialized instrumentation for XRF...