Recent studies have shown that reducing the precision of floating‐point calculations in an atmospheric model can improve the model's computational performance without affecting model fidelity, but code changes are needed to accommodate lower precision or to prevent undue round‐off error. For complex...
Filter results
Category
- (-) Computational Research (15)
- Scientific Discovery (248)
- Biology (153)
- Earth System Science (117)
- Human Health (92)
- Integrative Omics (59)
- Microbiome Science (26)
- National Security (20)
- Computing & Analytics (14)
- Energy Resiliency (9)
- Data Analytics & Machine Learning (7)
- Visual Analytics (6)
- Materials Science (5)
- Atmospheric Science (4)
- Chemical & Biological Signatures Science (4)
- Coastal Science (4)
- Renewable Energy (4)
- Weapons of Mass Effect (4)
- Chemistry (3)
- Computational Mathematics & Statistics (3)
- Data Analytics & Machine Learning (3)
- Cybersecurity (2)
- Distribution (2)
- Ecosystem Science (2)
- Electric Grid Modernization (2)
- Energy Efficiency (2)
- Energy Storage (2)
- Grid Cybersecurity (2)
- Plant Science (2)
- Solar Energy (2)
- Bioenergy Technologies (1)
- Computational Mathematics & Statistics (1)
- Grid Analytics (1)
- High-Performance Computing (1)
- Subsurface Science (1)
- Terrestrial Aquatics (1)
- Transportation (1)
- Wind Energy (1)
Publication Type
Tags
- Type 1 Diabetes (6)
- Autoimmunity (5)
- Machine Learning (5)
- Biomarkers (4)
- Molecular Profiling (4)
- Mass spectrometry-based Omics (3)
- Predictive Modeling (3)
- Alternative Splicing (2)
- DNA Sequence Analysis (1)
- Fish Detection (1)
- Functional Annotation Analysis (1)
- High-Performance Computing (1)
- Hydropower (1)
- Kmers (1)
- Marine and Hydrokinetic (1)
- Python (1)
- RNA Sequence Analysis (1)
- Snakemake (1)
- Software Data Analysis (1)
- underwater video (1)
Category
Christine H Chang, William C Nelson, Abby Jerger, Aaron T Wright, Robert G Egbert, Jason E McDermott, Snekmer: a scalable pipeline for protein sequence fingerprinting based on amino acid recoding, Bioinformatics Advances , Volume 3, Issue 1, 2023, vbad005, https://doi.org/10.1093/bioadv/vbad005...
The Human Islet Research Network (HIRN) is a large consortia with many research projects focused on understanding how beta cells are lost in type 1 diabetics (T1D) with a goal of finding how to protect against or replace the loss of functional beta cells. The consortia has multiple branches of...
Datasets
0
This project is an interdisciplinary collaboration supported by US DOE Office of Science's Scientific Discovery through Advanced Computing (SciDAC) program. The project addresses a crucial but largely overlooked source of error in the Energy Exascale Earth System Model (E3SM) and other atmosphere...
Category
Datasets
2
Category
Datasets
1
Machine learning is a core technology that is rapidly advancing within type 1 diabetes (T1D) research. Our Human Islet Research Network (HIRN) grant is studying early cellular response initiating β cell stress in T1D through the generation of heterogenous low- and high-throughput molecular...
Datasets
3
The Environmental Determinants of Diabetes in the Young (TEDDY) study is searching for factors influencing the development of type 1 diabetes (T1D) in children. Research has shown that there are certain genes that correlate to higher risk of developing T1D, but not all children with these genes...
Datasets
1
The Diabetes Autoimmunity Study in the Young (DAISY) seeks to find environmental factors that can trigger the development of type 1 diabetes (T1D) in children. DAISY follows children with high-risk of developing T1D based on family history or genetic markers. Genes, diets, infections, and...
Datasets
1
Category
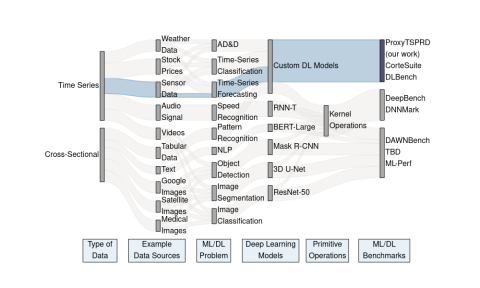
ProxyTSPRD profiles are collected using NVIDIA Nsight Systems version 2020.3.2.6-87e152c and capture computational patterns from training deep learning-based time-series proxy-applications on four different levels: models (Long short-term Memory and Convolutional Neural Network), DL frameworks...
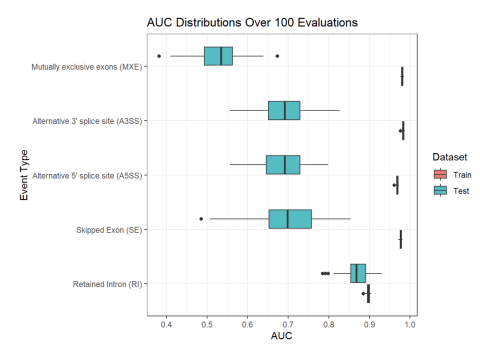
Inclusion levels of alternative splicing (AS) events of five different varieties (i.e. skipped exon (SE), retained intron (RI), alternative 5’ splice site (A5SS), alternative 3’ splice site (A3SS), and mutually exclusive exons (MXE)) were measured in human blood samples from two separate cohorts of...
A total of 172 children from the DAISY study with multiple plasma samples collected over time, with up to 23 years of follow-up, were characterized via proteomics analysis. Of the children there were 40 controls and 132 cases. All 132 cases had measurements across time relative to IA. Sampling was...
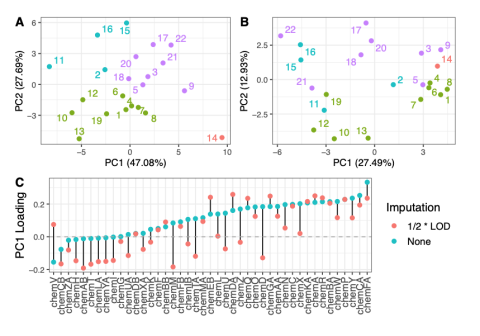
This data is supplementary to the manuscript Expanding the access of wearable silicone wristbands in community-engaged research through best practices in data analysis and integration by Lisa M. Bramer, Holly M. Dixon, David J. Degnan, Diana Rohlman, Julie B. Herbstman, Kim A. Anderson, and Katrina...
Comprised of 6,426 sample runs, The Environmental Determinants of Diabetes in the Young (TEDDY) proteomics validation study constitutes one of the largest targeted proteomics studies in the literature to date. Making quality control (QC) and donor sample data available to researchers aligns with...