Please cite as : Anderson L.N., W.C. Nelson, J.E. McDermott, R. Wu, S.J. Fansler, Y. Farris, and J.K. Jansson, et al. 2020. WA-TmG.1.0 (Metagenome, WA). [Data Set] PNNL DataHub. https://doi.org/10.25584/WATmG1/1635002 To enable a comprehensive survey of the metabolic potential of complex soil...
Filter results
Category
- (-) Earth System Science (5)
- (-) Computational Mathematics & Statistics (1)
- Scientific Discovery (8)
- Biology (7)
- Computational Research (2)
- Data Analytics & Machine Learning (2)
- Human Health (2)
- Computational Mathematics & Statistics (1)
- Computing & Analytics (1)
- Data Analytics & Machine Learning (1)
- Ecosystem Science (1)
- Microbiome Science (1)
- National Security (1)
Content type
Tags
- (-) Microbiome (4)
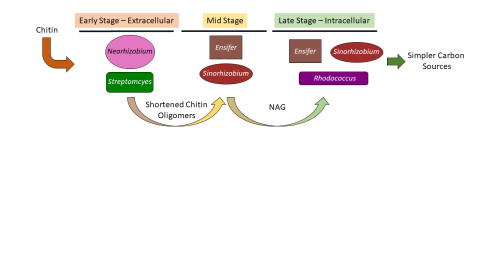
- (-) Chitin (1)
- (-) Machine Learning (1)
- Soil Microbiology (13)
- sequencing (9)
- Metagenomics (7)
- Omics (4)
- Fungi (3)
- Genomics (3)
- IAREC (3)
- metagenomics (3)
- Sequencing (3)
- soil microbiology (3)
- Climate Change (2)
- Mass Spectrometry (2)
- omics (2)
- Amplicon Sequencing (1)
- High Throughput Sequencing (1)
- Metatranscriptomic (1)
- microbiome stability (1)
- Microscopy (1)
- mineral weathering (1)
- MPLEX (1)
- MultiSector Dynamics (1)
- Output Databases (1)
- Software Data Analysis (1)
- Soil (1)
- species volatility (1)
- Spectroscopy (1)
- Viruses (1)
Category
Category
Please cite as : Anderson L.N., J.E. McDermott, and R.S. McClure. 2020. WA-IsoC_MSC1.1.0 (Amplicon 16S rRNA, WA). [Data Set] PNNL DataHub. https://doi.org/10.25584/WAIsoCMSC1/1635272 The soil microbiome is central to the cycling of carbon and other nutrients and to the promotion of plant growth...
Category
Please cite as : McClure R.S., Y. Farris, R.E. Danczak, W.C. Nelson, H. Song, A. Kessler, and J. Lee, et al. 2022. Metatranscriptomic data from MSC-2. [Data Set] PNNL DataHub. https://data.pnnl.gov/group/nodes/dataset/33232 Metatranscriptomic data from MSC-2 12 fastq files (6 forward read, 6 reverse...
Category
Comprised of 6,426 sample runs, The Environmental Determinants of Diabetes in the Young (TEDDY) proteomics validation study constitutes one of the largest targeted proteomics studies in the literature to date. Making quality control (QC) and donor sample data available to researchers aligns with...