Filter results
Content type
Tags
- (-) Microbiome (4)
- (-) Type 1 Diabetes (2)
- Soil Microbiology (13)
- sequencing (9)
- Metagenomics (7)
- Omics (4)
- Fungi (3)
- Genomics (3)
- IAREC (3)
- metagenomics (3)
- Sequencing (3)
- soil microbiology (3)
- Climate Change (2)
- Machine Learning (2)
- Mass Spectrometry (2)
- omics (2)
- Amplicon Sequencing (1)
- Bacterial Persistence (1)
- Bacterial Signaling (1)
- High Throughput Sequencing (1)
- microbiome stability (1)
- Microscopy (1)
- mineral weathering (1)
- MPLEX (1)
- MultiSector Dynamics (1)
- Output Databases (1)
- Software Data Analysis (1)
- species volatility (1)
- Spectroscopy (1)
- Viruses (1)
Please cite as : Anderson L.N., W.C. Nelson, J.E. McDermott, R. Wu, S.J. Fansler, Y. Farris, and J.K. Jansson, et al. 2020. WA-TmG.1.0 (Metagenome, WA). [Data Set] PNNL DataHub. https://doi.org/10.25584/WATmG1/1635002 To enable a comprehensive survey of the metabolic potential of complex soil...
Category
Category
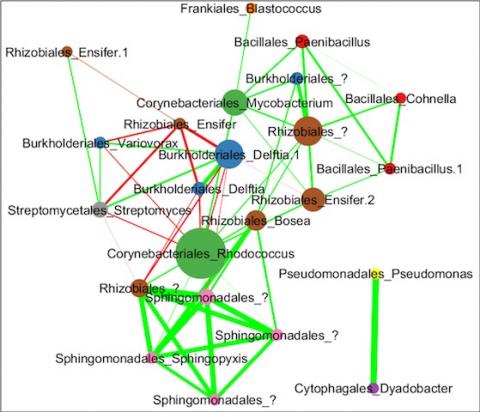
Please cite as : Anderson L.N., J.E. McDermott, and R.S. McClure. 2020. WA-IsoC_MSC1.1.0 (Amplicon 16S rRNA, WA). [Data Set] PNNL DataHub. https://doi.org/10.25584/WAIsoCMSC1/1635272 The soil microbiome is central to the cycling of carbon and other nutrients and to the promotion of plant growth...
Category
A total of 172 children from the DAISY study with multiple plasma samples collected over time, with up to 23 years of follow-up, were characterized via proteomics analysis. Of the children there were 40 controls and 132 cases. All 132 cases had measurements across time relative to IA. Sampling was...
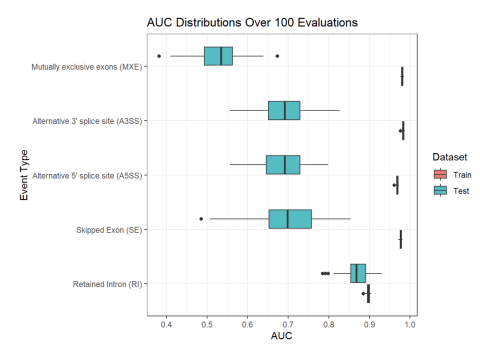
Inclusion levels of alternative splicing (AS) events of five different varieties (i.e. skipped exon (SE), retained intron (RI), alternative 5’ splice site (A5SS), alternative 3’ splice site (A3SS), and mutually exclusive exons (MXE)) were measured in human blood samples from two separate cohorts of...