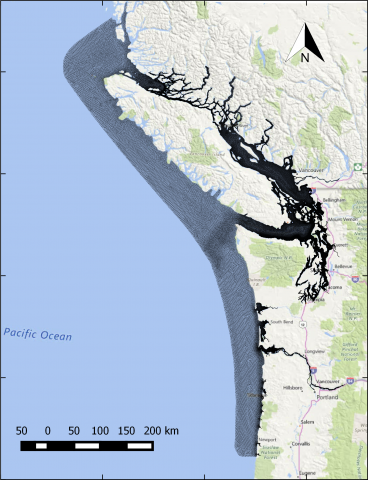
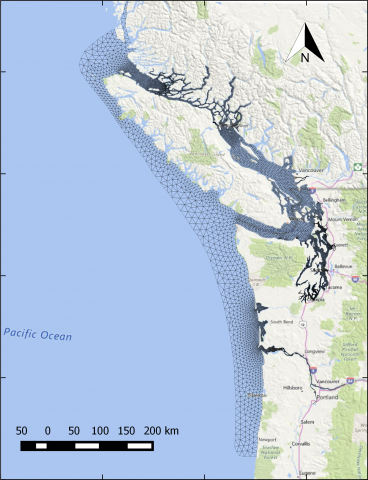
The year 2014 solution files are based on model calibration effort based on inputs and data from Year 2014 as described in Khangaonkar et al. 2018). It includes water surface elevation, currents, temperature and salinity at an hourly interval. These netcdf files with 24 hourly records include...
Filter results
Category
- (-) Scientific Discovery (211)
- (-) Grid Analytics (1)
- Biology (127)
- Earth System Science (93)
- Human Health (71)
- Integrative Omics (57)
- Microbiome Science (25)
- National Security (15)
- Computational Research (10)
- Computing & Analytics (10)
- Chemistry (7)
- Energy Resiliency (7)
- Visual Analytics (6)
- Computational Mathematics & Statistics (4)
- Data Analytics & Machine Learning (4)
- Atmospheric Science (3)
- Chemical & Biological Signatures Science (3)
- Coastal Science (3)
- Data Analytics & Machine Learning (3)
- Ecosystem Science (3)
- Renewable Energy (3)
- Weapons of Mass Effect (3)
- Cybersecurity (2)
- Distribution (2)
- Electric Grid Modernization (2)
- Grid Cybersecurity (2)
- Plant Science (2)
- Bioenergy Technologies (1)
- Computational Mathematics & Statistics (1)
- Energy Efficiency (1)
- Energy Storage (1)
- High-Performance Computing (1)
- Solar Energy (1)
- Subsurface Science (1)
- Terrestrial Aquatics (1)
- Transportation (1)
- Wind Energy (1)
Dataset Type
- Omics (83)
- Transcriptomics (51)
- Proteomics (38)
- Sequencing (38)
- Metabolomics (27)
- Lipidomics (26)
- Amplicon (16S, ITS) (14)
- Metagenomics (9)
- Microarray (9)
- Simulation Results (6)
- Genomics (4)
- Computed Analysis (3)
- ChIP-seq (2)
- Cybersecurity (1)
- Geospacial (1)
- Mass Spectra (1)
- Phenomics (1)
- Staff (1)
- Whole genome sequencing (WGS) (1)
Tags
- Virology (64)
- Immune Response (48)
- Time Sampled Measurement Datasets (45)
- Gene expression profile data (42)
- Differential Expression Analysis (41)
- Homo sapiens (30)
- Mass spectrometry data (25)
- Multi-Omics (25)
- MERS-CoV (16)
- Mus musculus (15)
- Viruses (15)
- Health (14)
- Virus (14)
- Soil Microbiology (13)
- West Nile virus (11)
- Ebola (9)
- Influenza A (9)
- sequencing (9)
- Resource Metadata (8)
- Mass Spectrometry (7)
- Metagenomics (7)
- Microarray (5)
- Omics (5)
- PerCon SFA (5)
- Genomics (4)
- Human Interferon (4)
- Microbiome (4)
- Proteomics (4)
- Sequencing (3)
- soil microbiology (3)
This data set provides ITS fungal community composition via DNA sequence analysis at the time of peat coring at the South End bog in 2013. These samples were collected outside the experimental enclosures and are pre-treatment with no experimental manipulation. These are part of the Spruce and...
Category
The year 2014 solution files are based on model calibration effort based on inputs and data from Year 2014 as described in Khangaonkar et al. 2018). It includes water surface elevation, currents, temperature and salinity at an hourly interval. These netcdf files with 24 hourly records include...
Category
This data set provides the ITS fungal community composition via DNA sequence analysis from sand and peat ingrowth cores at the South End bog in 2013. These samples were collected outside the experimental enclosures and are pre-treatment with no experimental manipulation. These are part of the Spruce...
Category
This data set provides the peat microbial biomass carbon (MBC) and nitrogen (MBN), extractable organic carbon (EOC) and extractable nitrogen (EN) at the time of peat coring for Deep Peat Heating (DPH) and Whole Ecosystem Warming (WEW) for 2014-2017 from the Spruce and Peatland Responses Under...
Category
This data set provides the ITS fungal community composition via DNA and cDNA sequence analysis for peat and sand ingrowth cores collected before and during Deep Peat Heating (DPH) and Whole Ecosystem Warming (WEW) for 2015-2016 from the Spruce and Peatlands Under Changing Environments (SPRUCE)...
Category
This data set provides the 16S microbial community composition via DNA sequence analysis at the time of peat coring at the South End bog in 2013. These samples were collected outside the experimental enclosures and are pre-treatment with no experimental manipulation. These are part of the Spruce and...
Category
This data set provides the ingrowth peat extracellular enzyme potential (EE) for before and during Deep Peat Heating (DPH) and Whole Ecosystem Warming (WEW) for 2015-2016 from the Spruce and Peatlands Under Changing Environments (SPRUCE). EE potential was quantified and calculated following a...
Category
This data set provides the 16S microbial community composition via DNA and cDNA sequence analyses at the time of peat coring for Deep Peat Heating (DPH) and Whole Ecosystem Warming (WEW) for 2014-2017 from the Spruce and Peatlands Under Changing Environments (SPRUCE). Samples were extracted using a...
Category
This data set provides the ingrowth peat microbial biomass carbon (MBC) and nitrogen (MBN), extractable organic carbon (EOC) and extractable nitrogen (EN) for Deep Peat Heating (DPH) and Whole Ecosystem Warming (WEW) for 2015-2016 from the Spruce and Peatland Responses Under Changing Environments...
Category
This data set provides the ingrowth peat extracellular enzyme potential (EE) for before and during Deep Peat Heating (DPH) and Whole Ecosystem Warming (WEW) for 2015-2016 from the Spruce and Peatlands Under Changing Environments (SPRUCE). EE potential was quantified and calculated following a...
Category
This data set provides the 16S microbial community composition of peat and sand ingrowth cores via DNA and cDNA sequence analysis before and during Deep Peat Heating (DPH) and Whole Ecosystem Warming (WEW) for 2015-2016 from the Spruce and Peatlands Under Changing Environments (SPRUCE). Samples were...
Category
This data set provides ITS fungal community composition via DNA and cDNA sequence analysis at the time of peat coring for Deep Peat Heating (DPH) and Whole Ecosystem Warming (WEW) for 2014-2017 from the Spruce and Peatlands Under Changing Environments (SPRUCE). Samples were extracted using a Qiagen...
Category
Please cite as : McClure R.S., Y. Farris, R.E. Danczak, W.C. Nelson, H. Song, A. Kessler, and J. Lee, et al. 2022. Model Soil Consortium 2 (MSC-2) Bacterial Isolate Genomes. [Data Set] PNNL DataHub. https://doi.org/10.25584/PNNLDH/1986536 Model Soil Consortium 2 (MSC-2) Bacterial Isolate Genomes...
Category
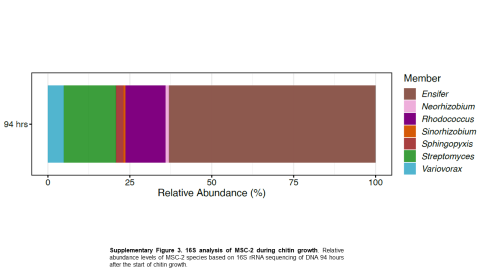
Please cite as : McClure R.S., Y. Farris, R.E. Danczak, W.C. Nelson, H. Song, A. Kessler, and J. Lee, et al. 2022. 16s data from MSC-2 growth. [Data Set] PNNL DataHub. https://data.pnnl.gov/group/nodes/dataset/33231 16s data from MSC-2 growth 3 fastq of 16s amplicon data of MSC2 1 csv file of raw...