Christine H Chang, William C Nelson, Abby Jerger, Aaron T Wright, Robert G Egbert, Jason E McDermott, Snekmer: a scalable pipeline for protein sequence fingerprinting based on amino acid recoding, Bioinformatics Advances , Volume 3, Issue 1, 2023, vbad005, https://doi.org/10.1093/bioadv/vbad005...
Filter results
Content type
Tags
- (-) DNA Sequence Analysis (2)
- (-) Output Databases (1)
- Machine Learning (6)
- Type 1 Diabetes (6)
- Autoimmunity (5)
- Synthetic (5)
- Biomarkers (4)
- Mass Spectrometry (4)
- Molecular Profiling (4)
- Predictive Modeling (4)
- Mass spectrometry-based Omics (3)
- Omics (3)
- Proteomics (3)
- Software Data Analysis (3)
- Alternative Splicing (2)
- Cybersecurity (2)
- Electrical energy (2)
- Functional Annotation Analysis (2)
- Kmers (2)
- Python (2)
- RNA Sequence Analysis (2)
- Snakemake (2)
- EBC (1)
- Exhaled Breath Condensate (1)
- High-Performance Computing (1)
- Mass Spectrometer (1)
- Quantification (1)
- Statistics (1)
- TMT (1)
- underwater video (1)
Fusarium sp. DS682 Proteogenomics Statistical Data Analysis of SFA dataset download: 10.25584/KSOmicsFspDS682/1766303 . GitHub Repository Source: https://github.com/lmbramer/Fusarium-sp.-DS-682-Proteogenomics MaxQuant Export Files (txt) Trelliscope Boxplots (jsonp) Fusarium Report (.Rmd, html)...
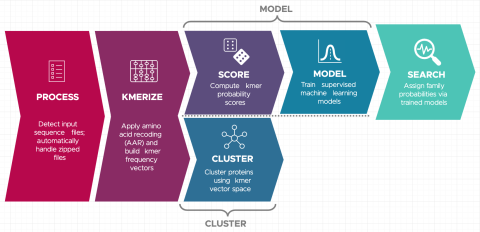
Last updated on 2023-02-23T19:37:46+00:00 by LN Anderson Snekmer: A scalable pipeline for protein sequence fingerprinting using amino acid recoding (AAR) Snekmer is a software package designed to reduce the representation of protein sequences by combining amino acid reduction (AAR) with the kmer...
Biography Young-Mo Kim is a senior bioanalytical chemist at PNNL. He received his PhD from Pohang University of Science and Technology in the School of Environmental Science and Engineering, studying analysis of metabolites during the bacterial and fungal degradation of xenobiotic substrates using...