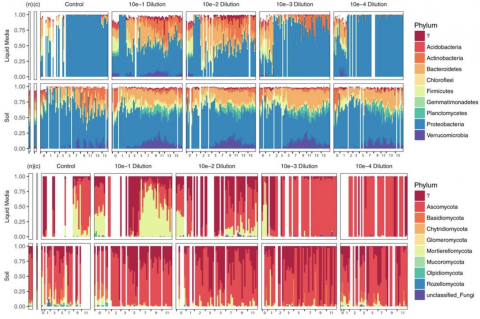
Please cite as : Zegeye E., C.J. Brislawn, Y. Farris, S.J. Fansler, K.S. Hofmockel, J.K. Jansson, and A.T. Wright, et al. 2019. WA-IsoC_NAG.1.0 (Amplicon 16S/ITS, WA). [Data Set] PNNL DataHub. https://dx.doi.org/10.25584/data.2019-02.700/1506698 Investigation of the successional dynamics of a soil...
Filter results
Content type
Tags
- (-) Soil Microbiology (3)
- (-) IAREC (1)
- Virology (26)
- Differential Expression Analysis (23)
- Immune Response (23)
- Mass spectrometry data (23)
- Time Sampled Measurement Datasets (23)
- Multi-Omics (22)
- Gene expression profile data (17)
- Homo sapiens (16)
- MERS-CoV (12)
- Mus musculus (7)
- Influenza A (5)
- West Nile virus (4)
- Health (3)
- Virus (3)
- Viruses (3)
- Ebola (2)
- sequencing (2)
- Fungi (1)
- High-Performance Computing (1)
- Mass Spectrometry (1)
- Microbiome (1)
- microbiome stability (1)
- MPLEX (1)
- MultiSector Dynamics (1)
- Omics (1)
- Output Databases (1)
- Software Data Analysis (1)
- species volatility (1)
Showing 1 - 3 of 3
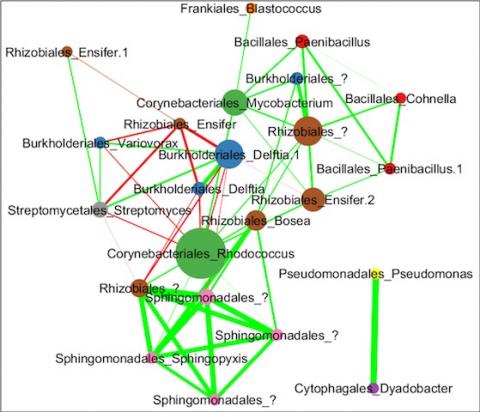
Please cite as : Anderson L.N., J.E. McDermott, and R.S. McClure. 2020. WA-IsoC_MSC1.1.0 (Amplicon 16S rRNA, WA). [Data Set] PNNL DataHub. https://doi.org/10.25584/WAIsoCMSC1/1635272 The soil microbiome is central to the cycling of carbon and other nutrients and to the promotion of plant growth...
Category
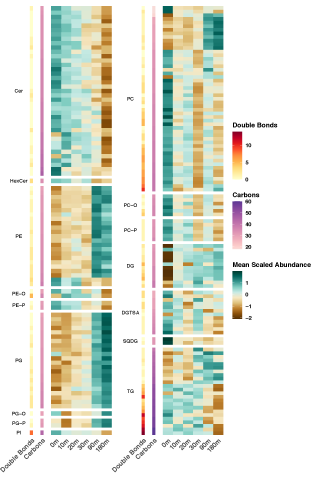
Rapid remodeling of the soil lipidome in response to a drying-rewetting event - Multi-Omics Data Package DOI Data package contents reported here are the first version and contain pre- and post-processed data acquisition and subsequent downstream analysis files using various data source instrument...