Category
Description
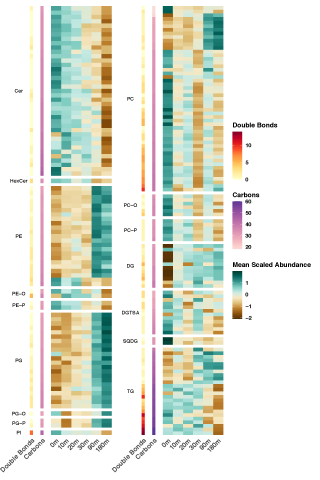
Rapid remodeling of the soil lipidome in response to a drying-rewetting event - Multi-Omics Data Package DOI
Data package contents reported here are the first version and contain pre- and post-processed data acquisition and subsequent downstream analysis files using various data source instrument method techniques and Mass Spectroscopy (MS) EMSL capabilities. This publication data package DOI is a comprehensive high-throughput multi-omics data lifecycle collection containing processed data method metadata. Support files include additional data download “Read Me” file containing data descriptor information and data source application ontologies (see data dictionary). Reported data download contents are structured for compliance with project data sharing guidelines, community standards initiatives, and sponsor stakeholder policies supporting FAIR data principles. For increased data availability and interoperability, GC-MS/LC-MS mass spectrometry datasets (Thermo .raw) were deposited at the MassIVE database repository and can be accessed by the link provided below. Statistical data processing software, analysis tools, and data workflows are listed below corresponding to the host repository long-term location.
Content Contributors
- Lindsey N Anderson - Data Curation
- Mark Pleake - IT Resource Support
Linked Open Data
- Mass Spectrometry MassIVE Accession: MSV000086931 (90 experimental runs; 18.73 GB) | MassIVE DOI: 10.25345/C5K50H
- BioProject Accession: PRJNA886746 | SRA Accession: SRP400851 (60 experimental runs; 1.2 GB)
Available Data Downloads (12.3 GB)
- "Read Me" data package content files (doc, txt)
- Standard structured metadata files (xls, json, xml)
- SwissLipids species ontology identifiers (csv)
- Processed data analysis R markdown (html, Rmd)
Accessible File formats
- Raw measurement data (LC-MS: raw, cdf, mzML; Amplicon: fastq)
- Processed/Data processing files (rmd, html, fasta, txt, csv, xlsx, mzXML)
- Image object files (pdf)
- Linked repository metadata collection files (xml)
- Technical parameters/specification files (csv, xml)
- Molecular Network (cys, html)
Linked Software
- LIQUID LC-MS Analysis Software | 10.5281/zenodo.6459462
- Lipid Mini-On Software Tools | 10.5281/zenodo.1492803
- GNPS
- MZmine 2
- Cytoscape
- pmartR Omics Statistical Software | 10.5281/zenodo.6108667
- QIIME2 (v2021.4)
- SILVA Database (v138)
- UNITE Database (v8-10.05.2021)
Funding Acknowledgements
This research was supported by the Department of Energy Office of Biological and Environmental Research (BER) and is a contribution of the Scientific Focus Area ‘Phenotypic response of the soil microbiome to environmental perturbations’. PNNL is operated for the DOE by Battelle Memorial Institute under Contract DE-AC05-76RLO1830. A portion of this work was performed under the EMSL user project award 10.46936/cpcy.proj.2021.60161/60000437, utilizing instrument capabilities operating at the Environmental Molecular Sciences Laboratory, a DOE Office of Science User Facility sponsored by the Biological and Environmental Research program under Contract No. DE-AC05-76RL01830.
Data Reuse Citation Policy
When reusing provided linked project dataset DOIs listed at the PNNL DataHub institutional project data repository, we ask that users please cite the primary journal article publication and the dataset digital object identifier listed at the download page. Reference citations, where applicable, provide the necessary metadata information required to support, corroborate, verify, and otherwise determine the legitimacy of the research findings provided (data and code) from scholarly publications and corresponding project data releases.
Cite as
- Anderson, Lindsey N, Couvillion, Sneha P, Pleake, Mark M, Stephan, Eric G, and Hofmockel, Kirsten S. 2021. WA-Omics_LA.1.0 (Metabolomics, Lipidomics, & Amplicon 16S/ITS, WA) [Data Set] PNNL DataHub. https://doi.org/10.25584/WAOmicsLA1/1866788
- Couvillion, S.P., Danczak, R.E., Naylor, D. et al. Rapid remodeling of the soil lipidome in response to a drying-rewetting event. Microbiome 11, 34 (2023). https://doi.org/10.1186/s40168-022-01427-4
Last updated on 2023-03-21T16:45:38+00:00 by LN Anderson