Category
Method Abstract
Last updated on 2024-03-21T18:47:20+00:00 by LN Anderson
MPLEx Method Protocol for Integrative Single-Sample Processing for Subsequent Proteomic, Metabolomic, and Lipidomic Analyses
Method Source Contributions
- LN Anderson, Data Curation
The Metabolite, Protein and Lipid Extraction (MPLEx) method protocol is a robust sample processing technique applicable to a breadth of diverse sample types, including various cell cultures, environmental collections, and delicate tissues. The MPLEx method is comprised of an adapted solvent-based method, widely applied for extracting functional characterization information from complex cellular structures containing metabolites, lipids, and protein for analysis by mass spectrometry-based qualitative and quantitative measurements.
Explore PNNL DataHub projects utilizing data source acquisition methods related to published data from integrated research platforms using high-resolution mass spectrometry method technologies (see example). Instrument capabilities are linked to published project datasets, publications, analyses, and externally related database resources from a suite of cross-discipline data science domain investigations.
Reference Citations:
- Nakayasu ES, Nicora CD, Sims AC, Burnum-Johnson KE, Kim YM, Kyle JE, Matzke MM, Shukla AK, Chu RK, Schepmoes AA, Jacobs JM, Baric RS, Webb-Robertson BJ, Smith RD, Metz TO. MPLEx: a Robust and Universal Protocol for Single-Sample Integrative Proteomic, Metabolomic, and Lipidomic Analyses. mSystems. 2016 May 10;1(3):e00043-16. doi: 10.1128/mSystems.00043-16.
- Nicora, C. D., Burnum-Johnson, K. E., Nakayasu, E. S., Casey, C. P., White III, R. A., Roy Chowdhury, T., Kyle, J. E., Kim, Y., Smith, R. D., Metz, T. O., Jansson, J. K., Baker, E. S. The MPLEx Protocol for Multi-omic Analyses of Soil Samples. JoVE (135), e57343, doi:10.3791/57343 (2018).
Watch the MPLEx video on JoVE
MPLEx Experimental Method Ontology
Laboratory Techniques: Experimental Protocol > Standard Operating Procedure > Mass Spectrometry Experimental Protocol
Data Acquisition Type: Mass Spectrometry
Topic Areas: Omics > Proteomics, Metabolomics, Lipidomics, Phenomics
For more information about complimentary downstream instrument capabilities leveraging the MPLEx sample preparation method, visit the EMSL instrument & resources homepage.
Acknowledgment of Federal Funding
The data described here was funded in whole or in part by the National Institute of Allergy and Infectious Diseases, of the National Institutes of Health under award number U19AI106772 and is a contribution of the "Modeling Host Responses to Understand Severe Human Virus Infections" Project at Pacific Northwest National Laboratory. Data generated by the Omics-LHV Core for proteomics, metabolomics, and lipidomics analyses for were performed at Pacific Northwest National Laboratory in the Environmental Molecular Sciences Laboratory, a national scientific user facility sponsored by the Department of Energy’s (DOE) Office, operating under the Battelle Memorial Institute for the DOE under contract number DE-AC05-76RLO1830.
Federal Acknowledgements
This work was supported in part by the Earth and Biological Sciences Directorate (EBSD) at Pacific Northwest National Laboratory (PNNL), a multiprogram national laboratory managed by the Battelle Memorial Institute, operating under the U.S. Department of Energy (DOE) contract DE-AC05-76RL01830. User capabilities described here reflect collaborations with the Environmental Molecular Sciences Laboratory (EMSL), a DOE Office of Science (SC-3) user facility operating under the Contract No. DE-AC05-76RL01830.
Terms of Use
Recommendation guidelines provided by the DOE Office of Science can be accessed at the SC Funding Opportunities & Acknowledgements homepage. For additional information regarding user capability data release, visit the SC Digital Data Management Resources at User Facilities for more information.
EMSL Acknowledgment
Scientists who publish results of research using Environmental Molecular Sciences Laboratory (EMSL) resources, capabilities, and resulting data are required to include an acknowledgment of EMSL Policies in any publications. Learn more about user facility data management resources at the Office of Science page.
Projects (7)
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson PNNL DataHub NIAID Program Project: Modeling Host Responses to Understand Severe Human Virus Infections, Multi-Omic Viral Dataset Catalog Collection Background The National Institute of Allergy and Infectious Diseases (NIAID) "Modeling Host...
Category
Datasets
45
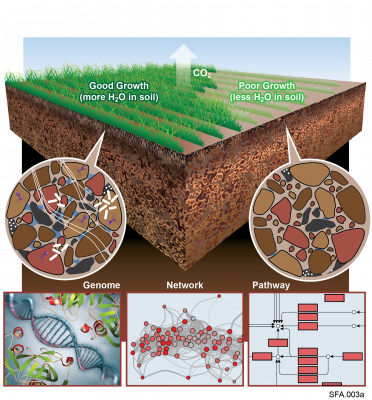
The Phenotypic Response of the Soil Microbiome to Environmental Perturbations Project (Soil Microbiome SFA) at Pacific Northwest National Laboratory is a Genomic Sciences Program Science Focus Area (SFA) Project operating under the Environmental Microbiome Science Research Area. The Soil Microbiome...
Datasets
45
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson Omics-LHV Profiling of Host Response to Ebola Virus Infection Background Ebola virus ( EBOV ) is a high risk biological agent, belonging to the Flaviviridae family, and is classified as a Category A priority pathogen by the National Institute...
Category
Datasets
6
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson Omics-LHV Profiling of Host Response to Influenza A Virus Infection Background Influenza A virus ( IAV ) is a high risk biological agent belonging to the Orthomyxoviridae family is classified as a Category C priority pathogen by the National...
Category
Datasets
8
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson Omics-LHV Profiling of Host Response to West Nile Virus Infection Background West Nile virus ( WNV ) belongs to the mosquito-borne Flaviviridae family and is classified as a Category A priority pathogen by the National Institute of Allergy and...
Category
Datasets
11
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson Omics-LHV Profiling of Host Response to MERS-CoV Virus Infection Background Middle East Respiratory Syndrome coronavirus ( MERS-CoV ), part of the Coronaviridae family, is classified as a Category C priority pathogen by the National Institute...
Category
Datasets
15
Last updated on 2023-02-23T19:37:46+00:00 by LN Anderson PerCon SFA Project Publication Experimental Data Catalog The Persistence Control of Engineered Functions in Complex Soil Microbiomes Project (PerCon SFA) at Pacific Northwest National Laboratory ( PNNL ) is a Genomic Sciences Program...
Datasets
5