This data set provides the peat microbial biomass carbon (MBC) and nitrogen (MBN), extractable organic carbon (EOC) and extractable nitrogen (EN) at the time of peat coring for Deep Peat Heating (DPH) and Whole Ecosystem Warming (WEW) for 2014-2017 from the Spruce and Peatland Responses Under...
Filter results
Category
- Biology (56)
- Scientific Discovery (56)
- Human Health (40)
- Integrative Omics (28)
- Microbiome Science (12)
- Earth System Science (4)
- Computational Mathematics & Statistics (2)
- Computational Research (2)
- Data Analytics & Machine Learning (2)
- Chemical & Biological Signatures Science (1)
- National Security (1)
- Weapons of Mass Effect (1)
Content type
Dataset Type
- (-) Proteomics (39)
- (-) Amplicon (16S, ITS) (15)
- (-) Microarray (9)
- Omics (86)
- Transcriptomics (51)
- Sequencing (39)
- Metabolomics (28)
- Lipidomics (26)
- Simulation Results (17)
- Metagenomics (9)
- Computed Analysis (4)
- Genomics (4)
- ChIP-seq (2)
- Geospacial (2)
- Staff (2)
- Cybersecurity (1)
- Electromagnetic Spectrum (1)
- Mass Spectra (1)
- Microscopy (1)
- Phenomics (1)
- Whole genome sequencing (WGS) (1)
Tags
- Virology (37)
- Mass spectrometry data (25)
- Differential Expression Analysis (24)
- Immune Response (24)
- Time Sampled Measurement Datasets (24)
- Multi-Omics (23)
- Gene expression profile data (18)
- Homo sapiens (16)
- MERS-CoV (13)
- Health (11)
- Virus (11)
- Viruses (11)
- Mus musculus (8)
- Influenza A (5)
- West Nile virus (5)
- Soil Microbiology (4)
- Ebola (2)
- Fungi (2)
- Mass Spectrometry (2)
- Omics (2)
- Proteomics (2)
- sequencing (2)
- Viral Experiment (2)
- EBC (1)
- IAREC (1)
- microbiome stability (1)
- Microscopy (1)
- Quantification (1)
- species volatility (1)
- TMT (1)
This data set provides the 16S microbial community composition via DNA sequence analysis at the time of peat coring at the South End bog in 2013. These samples were collected outside the experimental enclosures and are pre-treatment with no experimental manipulation. These are part of the Spruce and...
Category
This data set provides the 16S microbial community composition of peat and sand ingrowth cores via DNA and cDNA sequence analysis before and during Deep Peat Heating (DPH) and Whole Ecosystem Warming (WEW) for 2015-2016 from the Spruce and Peatlands Under Changing Environments (SPRUCE). Samples were...
Category
This data set provides ITS fungal community composition via DNA and cDNA sequence analysis at the time of peat coring for Deep Peat Heating (DPH) and Whole Ecosystem Warming (WEW) for 2014-2017 from the Spruce and Peatlands Under Changing Environments (SPRUCE). Samples were extracted using a Qiagen...
Category
This data set provides the ITS fungal community composition via DNA and cDNA sequence analysis for peat and sand ingrowth cores collected before and during Deep Peat Heating (DPH) and Whole Ecosystem Warming (WEW) for 2015-2016 from the Spruce and Peatlands Under Changing Environments (SPRUCE)...
Category
This data set provides the ingrowth peat extracellular enzyme potential (EE) for before and during Deep Peat Heating (DPH) and Whole Ecosystem Warming (WEW) for 2015-2016 from the Spruce and Peatlands Under Changing Environments (SPRUCE). EE potential was quantified and calculated following a...
Category
This data set provides the 16S microbial community composition via DNA and cDNA sequence analyses at the time of peat coring for Deep Peat Heating (DPH) and Whole Ecosystem Warming (WEW) for 2014-2017 from the Spruce and Peatlands Under Changing Environments (SPRUCE). Samples were extracted using a...
Category
This data set provides the ingrowth peat microbial biomass carbon (MBC) and nitrogen (MBN), extractable organic carbon (EOC) and extractable nitrogen (EN) for Deep Peat Heating (DPH) and Whole Ecosystem Warming (WEW) for 2015-2016 from the Spruce and Peatland Responses Under Changing Environments...
Category
A pre-trained convolutional neural network was fine-tuned for three separate classification tasks, distinguishing 2D images of: 1) single amino acids, 2) protein structural ball and stick images of metalloproteins, and 3) protein structural ball and stick images of metalloenzymes with the metal...
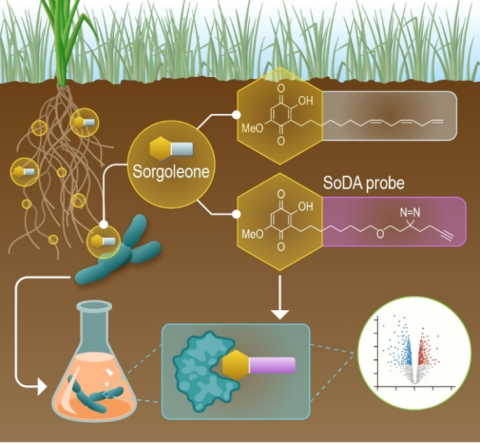
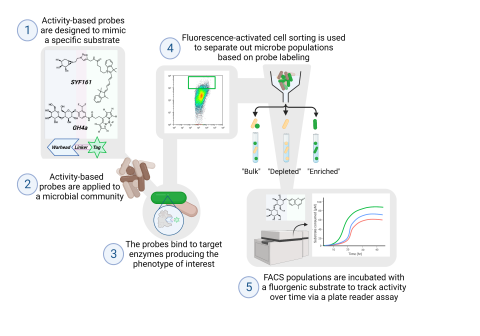
Last updated on 2024-04-19T19:12:08+00:00 by LN Anderson PerCon SFA: Profiling sorghum-microbe interactions with a specialized photoaffinity probe identifies key sorgoleone binders in Acinetobacter pitti Mass spectrometry-based global proteome analysis and SoDA-PAL photoaffinity probe labeled...
A total of 172 children from the DAISY study with multiple plasma samples collected over time, with up to 23 years of follow-up, were characterized via proteomics analysis. Of the children there were 40 controls and 132 cases. All 132 cases had measurements across time relative to IA. Sampling was...
16S amplicon fastq files to be associated with PhenoSelection manuscript.
Comprised of 6,426 sample runs, The Environmental Determinants of Diabetes in the Young (TEDDY) proteomics validation study constitutes one of the largest targeted proteomics studies in the literature to date. Making quality control (QC) and donor sample data available to researchers aligns with...