Christine H Chang, William C Nelson, Abby Jerger, Aaron T Wright, Robert G Egbert, Jason E McDermott, Snekmer: a scalable pipeline for protein sequence fingerprinting based on amino acid recoding, Bioinformatics Advances , Volume 3, Issue 1, 2023, vbad005, https://doi.org/10.1093/bioadv/vbad005...
Filter results
Category
- (-) Computational Research (4)
- (-) Integrative Omics (1)
- Scientific Discovery (31)
- Biology (29)
- Human Health (23)
- Earth System Science (4)
- Computational Mathematics & Statistics (1)
- Computing & Analytics (1)
- Data Analytics & Machine Learning (1)
- High-Performance Computing (1)
- Microbiome Science (1)
- National Security (1)
Content type
Tags
- Virology (57)
- Immune Response (53)
- Time Sampled Measurement Datasets (50)
- Gene expression profile data (47)
- Differential Expression Analysis (46)
- Homo sapiens (34)
- Mass spectrometry data (30)
- Multi-Omics (30)
- MERS-CoV (18)
- Mus musculus (18)
- West Nile virus (13)
- Ebola (11)
- Influenza A (11)
- Omics (10)
- Resource Metadata (9)
- Mass Spectrometry (8)
- Human Interferon (6)
- Microarray (6)
- Omics-LHV Project (6)
- Type 1 Diabetes (6)
- Autoimmunity (5)
- Machine Learning (5)
- Proteomics (5)
- Biomarkers (4)
- Mass Spectrometer (4)
- Mass spectrometry-based Omics (4)
- Molecular Profiling (4)
- Software Data Analysis (4)
- Imaging (3)
- Microscopy (3)
Omics-LHV, West Nile Experiment WGCN004 The purpose of this West Nile experiment was to obtain samples for omics analysis in primary mouse granule neuron cells infected with wild type West Nile virus (WNV-NY99 clone 382, WNVWT) and mutant virus (WNVE218A). Overall Design: Granule cell neurons from...
Category
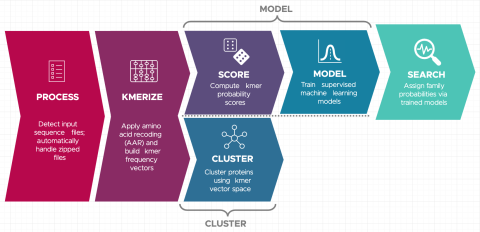
Last updated on 2023-02-23T19:37:46+00:00 by LN Anderson Snekmer: A scalable pipeline for protein sequence fingerprinting using amino acid recoding (AAR) Snekmer is a software package designed to reduce the representation of protein sequences by combining amino acid reduction (AAR) with the kmer...
Fusarium sp. DS682 Proteogenomics Statistical Data Analysis of SFA dataset download: 10.25584/KSOmicsFspDS682/1766303 . GitHub Repository Source: https://github.com/lmbramer/Fusarium-sp.-DS-682-Proteogenomics MaxQuant Export Files (txt) Trelliscope Boxplots (jsonp) Fusarium Report (.Rmd, html)...
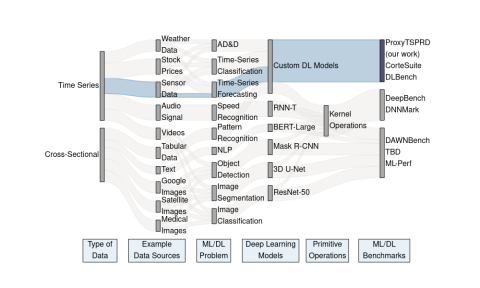
ProxyTSPRD profiles are collected using NVIDIA Nsight Systems version 2020.3.2.6-87e152c and capture computational patterns from training deep learning-based time-series proxy-applications on four different levels: models (Long short-term Memory and Convolutional Neural Network), DL frameworks...