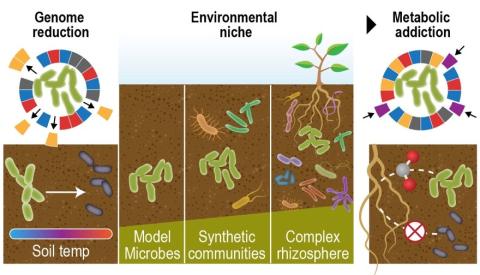
The rhizosphere represents a dynamic and complex interface between plant hosts and the microbial community found in the surrounding soil. While it is recognized that manipulating the rhizosphere has the potential to improve plant fitness and health, engineering the rhizosphere microbiome through...
Filter results
Category
- (-) Microbiome Science (4)
- (-) Computing & Analytics (1)
- Biology (10)
- Scientific Discovery (10)
- Computational Research (5)
- Data Analytics & Machine Learning (5)
- Human Health (5)
- Earth System Science (2)
- Computational Mathematics & Statistics (1)
- Computational Mathematics & Statistics (1)
- Data Analytics & Machine Learning (1)
- National Security (1)
Content type
Tags
- (-) Machine Learning (2)
- (-) Sorghum bicolor (2)
- (-) Whole Genome Sequencing (2)
- Omics (9)
- PerCon SFA (9)
- High Throughput Sequencing (8)
- Genomics (7)
- Sequencer System (5)
- Synthetic (5)
- Synthetic Biology (5)
- Mass Spectrometry (3)
- A. pittii SO1 (2)
- Amplicon Sequencing (2)
- Biological and Environmental Research (2)
- Imaging (2)
- Long Read Sequencer (2)
- Mass spectrometry data (2)
- Proteomics (2)
- RNA Sequence Analysis (2)
- Spectroscopy (2)
- Statistical Expression Analysis (2)
- Data Analysis (1)
- DOE (1)
- Environmental Microbiome Science (1)
- High-Performance Computing (1)
- Lipidomics (1)
- Mass Spectrometer (1)
- metabolomics (1)
- Output Databases (1)
- Soil Microbiology (1)
Metabolite exchange between plant roots and their associated rhizosphere microbiomes underpins plant growth promotion by microbes. Sorghum bicolor is a cereal crop that feeds animals and humans and is used for bioethanol production. Its root tips exude large amounts of a lipophilic benzoquinone...
A major challenge in biotechnology and biomanufacturing is the identification of a set of biomarkers for perturbations and metabolites of interest. Here, we develop a data-driven, transcriptome-wide approach to rank perturbation-inducible genes from time-series RNA sequencing data for the discovery...
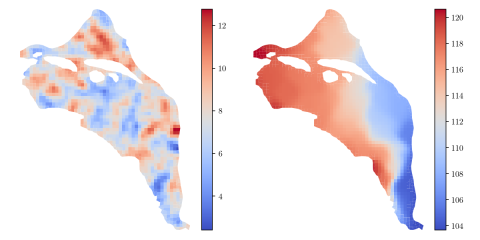
HDF5 file containing 10,000 hydraulic transmissivity inputs and the corresponding hydraulic pressure field outputs for a two-dimensional saturated flow model of the Hanford Site. The inputs are generated by sampling a 1,000-dimensional Kosambi-Karhunen-Loève (KKL) model of the transmissivity field...
Last updated on 2023-05-31T16:35:53+00:00 by LN Anderson PerCon SFA: Sequencing of Sorgoleone Promoting Rhizobacteria Isolates Whole genome sequencing (WGS) of sorgoleone utilizing rhizobacteria strains Pseudomonas sorgoleonovorans SO81 , Burkholderia anthina SO82 , and Acinetobacter pittii SO1 , as...