The srpAnalytics modeling pipeline for the Superfund Research Program Analytics Portal.
Filter results
Category
- (-) Biology (25)
- (-) Computational Mathematics & Statistics (2)
- Scientific Discovery (37)
- Microbiome Science (10)
- Chemistry (6)
- Earth System Science (6)
- Computational Research (5)
- Integrative Omics (5)
- Human Health (2)
- Chemical & Biological Signatures Science (1)
- Data Analytics & Machine Learning (1)
- Ecosystem Science (1)
- National Security (1)
- Plant Science (1)
- Weapons of Mass Effect (1)
Tags
- Mass Spectrometry (5)
- Multi-Omics (2)
- PerCon SFA (2)
- Proteomics (2)
- Synthetic Biology (2)
- Activity-Based Protein Profiling (1)
- Bacterial Persistence (1)
- Bacterial Signaling (1)
- Bioenergy Production (1)
- Chemical Biology (1)
- Chemical Probes (1)
- Data Analysis (1)
- Enzymology (1)
- Experimental Techniques (1)
- Functional Genomics (1)
- Health (1)
- Instrument Development (1)
- Metabolic Engineering (1)
- Metabolic Networks (1)
- metabolomics (1)
- Microbial Physiology (1)
- Microbiology (1)
- Omics (1)
- Output Databases (1)
- Protein Engineering (1)
- Structures for Lossless Ion Manipulations (1)
- Synthetic Chemistry (1)
- Virology (1)
- Virus (1)
- Viruses (1)
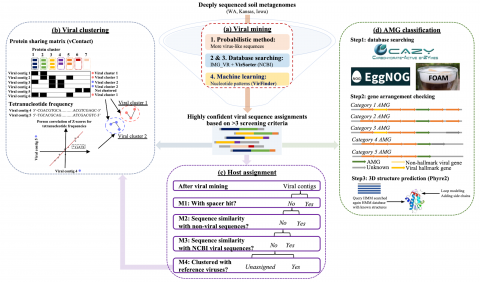
Code pertaining to the Soil Microbiome SFA Project publication data visualizations 'DNA viral diversity, abundance and functional potential vary across grassland soils with a range of historical moisture regimes' for processing publication data downloads.
Category
Data Science & Biostatistics
Category
Dr. Paul Piehowski is the Proteomics team leader for PNNL’s Environmental and Molecular Sciences Division and the Environmental Molecular Sciences Laboratory (EMSL) user program. Piehowski is an analytical chemist whose research is focused on the application of mass spectrometry to biological...
Category
The following R source code was used for plotting figures of the viral communities detected from three grasslands soil metagenomes with a historical precipitation gradient ( WA-TmG.2.0 , KS-TmG.2.0 , IA-TmG.2.0 ) from project publication 'DNA viral diversity, abundance and functional potential vary...
Category
Fusarium sp. DS682 Proteogenomics Statistical Data Analysis of SFA dataset download: 10.25584/KSOmicsFspDS682/1766303 . GitHub Repository Source: https://github.com/lmbramer/Fusarium-sp.-DS-682-Proteogenomics MaxQuant Export Files (txt) Trelliscope Boxplots (jsonp) Fusarium Report (.Rmd, html)...
Category
Category
Category
Category
Short Biography Caroline (Carrie) Harwood received her Ph.D. in microbiology from the University of Massachusetts and completed postdoctoral work at Yale University. She held academic appointments at Cornell University and the University of Iowa before moving to the University of Washington in 2005...
Category
Dr. Joshua Elmore is a Chemical Engineer at Pacific Northwest National Laboratory. He received his Ph. D. from the University of Georgia in the Department of Molecular Biology and Biochemistry studying the mechanisms of CRISPR/Cas microbial immune systems in hyperthermophilic Archaea. He was a...
Category
Biography Ernesto Nakayasu is a senior research scientist focused on understanding molecular mechanisms of diseases. Nakayasu has been applying systems biology and mass spectrometry-based omics measurements to study how pathogens and metabolic alterations cause human diseases; the goal is to...
Category
PerCon SFA, Co-Investigator Vivian Lin earned her PhD in organic chemistry from the University of California, Berkeley with Professor Chris Chang, developing fluorescent probes for imaging redox active small molecules. Afterward, she traveled to Switzerland for a postdoctoral fellowship in the...