Filter results
Tags
- (-) Virology (8)
- (-) High Throughput Sequencing (2)
- (-) Kmers (2)
- Viruses (11)
- Soil Microbiology (9)
- Health (8)
- Virus (8)
- Omics (5)
- PerCon SFA (5)
- Mass Spectrometry (4)
- Microbiome (4)
- RNA Sequence Analysis (4)
- sequencing (4)
- Fungi (3)
- Metagenomics (3)
- Software Data Analysis (3)
- Synthetic Biology (3)
- DNA Sequence Analysis (2)
- Functional Annotation Analysis (2)
- Genomics (2)
- Lipidomics (2)
- microbiome stability (2)
- Python (2)
- Snakemake (2)
- Sorghum bicolor (2)
- microbiome (1)
- Sequencing (1)
- soil microbiology (1)
- species volatility (1)
- Statistics (1)
Category
Category
Category
Category
Category
Category
Category
Elmore JR, Dexter GN, Baldino H, Huenemann JD, Francis R, Peabody GL 5th, Martinez-Baird J, Riley LA, Simmons T, Coleman-Derr D, Guss AM, Egbert RG. High-throughput genetic engineering of nonmodel and undomesticated bacteria via iterative site-specific genome integration. Sci Adv. 2023 Mar 10;9(10)...
Christine H Chang, William C Nelson, Abby Jerger, Aaron T Wright, Robert G Egbert, Jason E McDermott, Snekmer: a scalable pipeline for protein sequence fingerprinting based on amino acid recoding, Bioinformatics Advances , Volume 3, Issue 1, 2023, vbad005, https://doi.org/10.1093/bioadv/vbad005...
Metabolite exchange between plant roots and their associated rhizosphere microbiomes underpins plant growth promotion by microbes. Sorghum bicolor is a cereal crop that feeds animals and humans and is used for bioethanol production. Its root tips exude large amounts of a lipophilic benzoquinone...
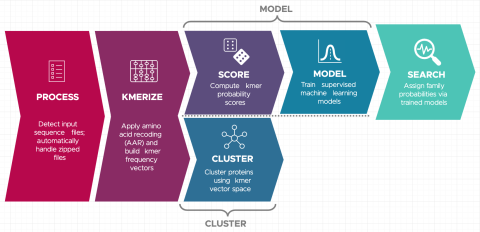
Last updated on 2023-02-23T19:37:46+00:00 by LN Anderson Snekmer: A scalable pipeline for protein sequence fingerprinting using amino acid recoding (AAR) Snekmer is a software package designed to reduce the representation of protein sequences by combining amino acid reduction (AAR) with the kmer...