Comprehensive assessment of climate datasets created by statistical or dynamical models is important for effectively communicating model projection and associated uncertainty to stakeholders and decision-makers. The Department of Energy FACETS project aims to foster such communication through...
Filter results
Category
- (-) Scientific Discovery (221)
- (-) Visual Analytics (6)
- Biology (136)
- Earth System Science (108)
- Human Health (80)
- Integrative Omics (52)
- Microbiome Science (23)
- National Security (15)
- Computing & Analytics (10)
- Computational Research (9)
- Energy Resiliency (7)
- Materials Science (5)
- Atmospheric Science (4)
- Chemical & Biological Signatures Science (3)
- Coastal Science (3)
- Computational Mathematics & Statistics (3)
- Data Analytics & Machine Learning (3)
- Data Analytics & Machine Learning (3)
- Renewable Energy (3)
- Weapons of Mass Effect (3)
- Chemistry (2)
- Cybersecurity (2)
- Distribution (2)
- Ecosystem Science (2)
- Electric Grid Modernization (2)
- Grid Cybersecurity (2)
- Plant Science (2)
- Bioenergy Technologies (1)
- Computational Mathematics & Statistics (1)
- Energy Efficiency (1)
- Energy Storage (1)
- Grid Analytics (1)
- High-Performance Computing (1)
- Solar Energy (1)
- Subsurface Science (1)
- Terrestrial Aquatics (1)
- Transportation (1)
- Wind Energy (1)
Dataset Type
- Omics (83)
- Transcriptomics (51)
- Proteomics (38)
- Sequencing (38)
- Metabolomics (27)
- Lipidomics (26)
- Amplicon (16S, ITS) (14)
- Metagenomics (9)
- Microarray (9)
- Simulation Results (6)
- Genomics (4)
- Computed Analysis (3)
- ChIP-seq (2)
- Staff (2)
- Geospacial (1)
- Mass Spectra (1)
- Phenomics (1)
- Whole genome sequencing (WGS) (1)
Publication Type
Tags
- Virology (71)
- Immune Response (48)
- Time Sampled Measurement Datasets (45)
- Gene expression profile data (42)
- Differential Expression Analysis (41)
- Homo sapiens (30)
- Mass spectrometry data (25)
- Viruses (24)
- Multi-Omics (23)
- Health (21)
- Soil Microbiology (21)
- Virus (21)
- MERS-CoV (16)
- Mus musculus (15)
- sequencing (13)
- West Nile virus (11)
- Ebola (9)
- Influenza A (9)
- Metagenomics (9)
- Microbiome (8)
- PerCon SFA (8)
- Resource Metadata (8)
- Omics (7)
- Fungi (6)
- Genomics (6)
- Microarray (6)
- Mass Spectrometry (4)
- metagenomics (4)
- Sequencing (4)
- soil microbiology (4)
Showing 136 - 150 of 227
Dataset
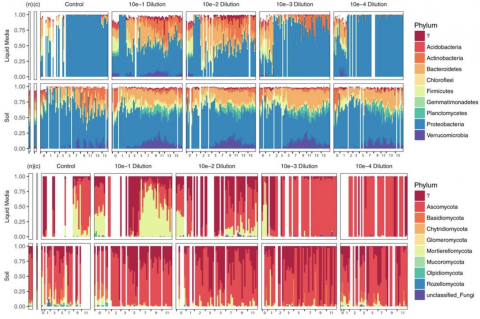
Please cite as : Anderson L.N., W.C. Nelson, J.E. McDermott, R. Wu, S.J. Fansler, Y. Farris, and J.K. Jansson, et al. 2020. IA-TmG.1.0 (Metagenome, IA). [Data Set] PNNL DataHub. https://doi.org/10.25584/IATmG1/1635005 To enable a comprehensive survey of the metabolic potential of complex soil...
Category
Dataset
Please cite as : Anderson L.N., W.C. Nelson, J.E. McDermott, R. Wu, S.J. Fansler, Y. Farris, and J.K. Jansson, et al. 2020. KS-TmG.1.0 (Metagenome, KS). [Data Set] PNNL DataHub. https://doi.org/10.25584/KSTmG1/1635004 To enable a comprehensive survey of the metabolic potential of complex soil...
Category
Dataset
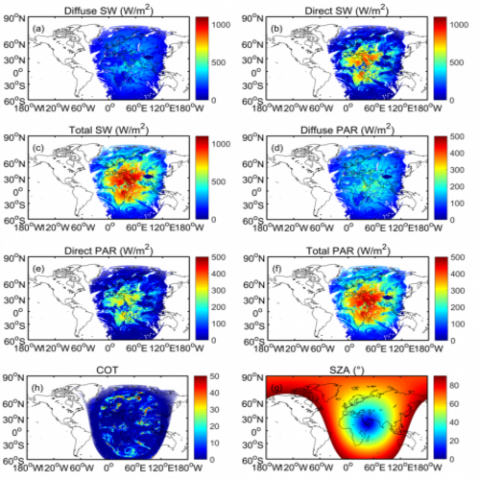
Both the hourly and daily data are provided in the product. The hourly data are grouped by day in distinct NetCDF files, which are named as “EPIC_SW_PAR_Hourly_yyyymmdd.nc” where “yyyy”, “mm”, and “dd” denote year, month, and day (UTC time). The daily data are grouped by month in distinct NetCDF...
Category
Dataset
Both the hourly and daily data are provided in the product. The hourly data are grouped by day in distinct NetCDF files, which are named as “EPIC_SW_PAR_Hourly_yyyymmdd.nc” where “yyyy”, “mm”, and “dd” denote year, month, and day (UTC time). The daily data are grouped by month in distinct NetCDF...
Category
Dataset
Both the hourly and daily data are provided in the product. The hourly data are grouped by day in distinct NetCDF files, which are named as “EPIC_SW_PAR_Hourly_yyyymmdd.nc” where “yyyy”, “mm”, and “dd” denote year, month, and day (UTC time). The daily data are grouped by month in distinct NetCDF...
Category
Dataset
Both the hourly and daily data are provided in the product. The hourly data are grouped by day in distinct NetCDF files, which are named as “EPIC_SW_PAR_Hourly_yyyymmdd.nc” where “yyyy”, “mm”, and “dd” denote year, month, and day (UTC time). The daily data are grouped by month in distinct NetCDF...
Category
Dataset
Both the hourly and daily data are provided in the product. The hourly data are grouped by day in distinct NetCDF files, which are named as “EPIC_SW_PAR_Hourly_yyyymmdd.nc” where “yyyy”, “mm”, and “dd” denote year, month, and day (UTC time). The daily data are grouped by month in distinct NetCDF...
Category
Dataset
Both the hourly and daily data are provided in the product. The hourly data are grouped by day in distinct NetCDF files, which are named as “EPIC_SW_PAR_Hourly_yyyymmdd.nc” where “yyyy”, “mm”, and “dd” denote year, month, and day (UTC time). The daily data are grouped by month in distinct NetCDF...
Category
Dataset
Both the hourly and daily data are provided in the product. The hourly data are grouped by day in distinct NetCDF files, which are named as “EPIC_SW_PAR_Hourly_yyyymmdd.nc” where “yyyy”, “mm”, and “dd” denote year, month, and day (UTC time). The daily data are grouped by month in distinct NetCDF...
Category
Dataset
Both the hourly and daily data are provided in the product. The hourly data are grouped by day in distinct NetCDF files, which are named as “EPIC_SW_PAR_Hourly_yyyymmdd.nc” where “yyyy”, “mm”, and “dd” denote year, month, and day (UTC time). The daily data are grouped by month in distinct NetCDF...
Category
Dataset
Please cite as : Anderson L.N., W.C. Nelson, J.E. McDermott, R. Wu, S.J. Fansler, Y. Farris, and J.K. Jansson, et al. 2020. WA-TmG.1.0 (Metagenome, WA). [Data Set] PNNL DataHub. https://doi.org/10.25584/WATmG1/1635002 To enable a comprehensive survey of the metabolic potential of complex soil...
Category
Please cite as : Zegeye E., C.J. Brislawn, Y. Farris, S.J. Fansler, K.S. Hofmockel, J.K. Jansson, and A.T. Wright, et al. 2019. WA-IsoC_NAG.1.0 (Amplicon 16S/ITS, WA). [Data Set] PNNL DataHub. https://dx.doi.org/10.25584/data.2019-02.700/1506698 Investigation of the successional dynamics of a soil...
Category
Dataset
Both the hourly and daily data are provided in the product. The hourly data are grouped by day in distinct NetCDF files, which are named as “EPIC_SW_PAR_Hourly_yyyymmdd.nc” where “yyyy”, “mm”, and “dd” denote year, month, and day (UTC time). The daily data are grouped by month in distinct NetCDF...
Category
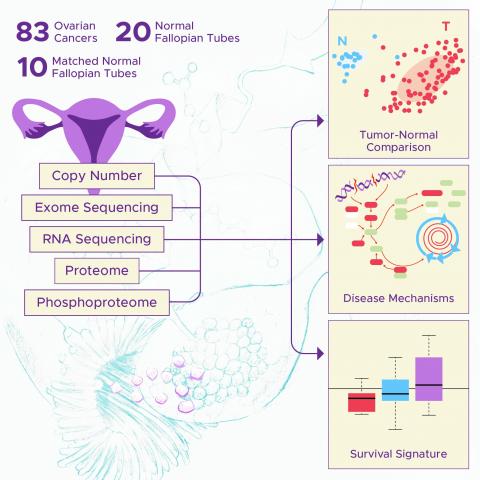
Interactive data plots for proteomics and phosphoproteomics data from tumor (n=83) and normal tissue from high-grade serous ovarian cancer.