"Visualizing the Hidden Half: Plant-Microbe Interactions in the Rhizosphere" Plant roots and the associated rhizosphere constitute a dynamic environment that fosters numerous intra- and interkingdom interactions, including metabolite exchange between plants and soil mediated by root exudates and the...
Filter results
Content type
Tags
- (-) Spectroscopy (2)
- (-) Whole Genome Sequencing (2)
- Omics (10)
- PerCon SFA (10)
- High Throughput Sequencing (8)
- Genomics (7)
- Mass Spectrometry (6)
- Synthetic Biology (6)
- Sequencer System (5)
- Proteomics (3)
- A. pittii SO1 (2)
- Amplicon Sequencing (2)
- Biological and Environmental Research (2)
- Cybersecurity (2)
- Electrical energy (2)
- Energy Equity (2)
- Energy Storage (2)
- Imaging (2)
- Long Read Sequencer (2)
- Mass Spectrometer (2)
- Mass spectrometry data (2)
- metabolomics (2)
- RNA Sequence Analysis (2)
- Sorghum bicolor (2)
- Statistical Expression Analysis (2)
- DOE (1)
- Lipidomics (1)
- Output Databases (1)
- Soil Microbiology (1)
- ToF-SIMS (1)
Metabolite exchange between plant roots and their associated rhizosphere microbiomes underpins plant growth promotion by microbes. Sorghum bicolor is a cereal crop that feeds animals and humans and is used for bioethanol production. Its root tips exude large amounts of a lipophilic benzoquinone...
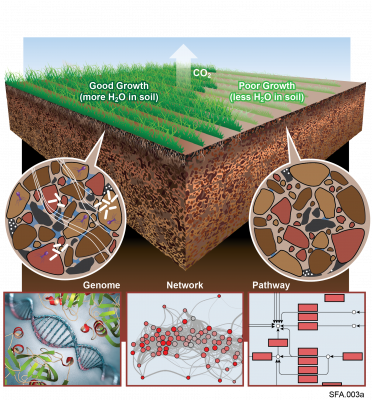
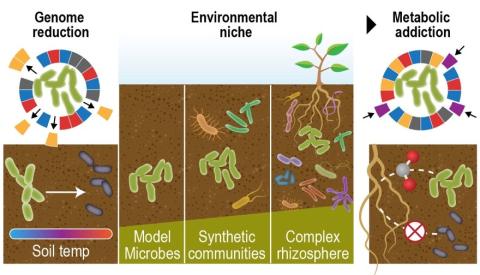
The Phenotypic Response of the Soil Microbiome to Environmental Perturbations Project (Soil Microbiome SFA) at Pacific Northwest National Laboratory is a Genomic Sciences Program Science Focus Area (SFA) Project operating under the Environmental Microbiome Science Research Area. The Soil Microbiome...
Datasets
23
Last updated on 2023-05-31T16:35:53+00:00 by LN Anderson PerCon SFA: Sequencing of Sorgoleone Promoting Rhizobacteria Isolates Whole genome sequencing (WGS) of sorgoleone utilizing rhizobacteria strains Pseudomonas sorgoleonovorans SO81 , Burkholderia anthina SO82 , and Acinetobacter pittii SO1 , as...