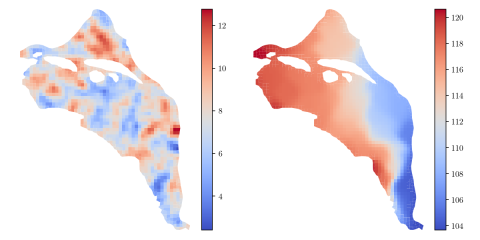
HDF5 file containing 10,000 hydraulic transmissivity inputs and the corresponding hydraulic pressure field outputs for a two-dimensional saturated flow model of the Hanford Site. The inputs are generated by sampling a 1,000-dimensional Kosambi-Karhunen-Loève (KKL) model of the transmissivity field...
Filter results
Category
- (-) Human Health (39)
- (-) National Security (2)
- Biology (41)
- Scientific Discovery (41)
- Integrative Omics (37)
- Computational Research (3)
- Data Analytics & Machine Learning (3)
- Chemical & Biological Signatures Science (1)
- Computational Mathematics & Statistics (1)
- Computing & Analytics (1)
- Data Analytics & Machine Learning (1)
- Weapons of Mass Effect (1)
Content type
Tags
- (-) Multi-Omics (28)
- (-) Mus musculus (18)
- (-) Predictive Modeling (4)
- Virology (79)
- Immune Response (53)
- Time Sampled Measurement Datasets (50)
- Gene expression profile data (47)
- Differential Expression Analysis (46)
- Homo sapiens (34)
- Mass spectrometry data (30)
- Health (23)
- Virus (23)
- Viruses (23)
- MERS-CoV (18)
- West Nile virus (13)
- Ebola (11)
- Influenza A (11)
- Resource Metadata (8)
- Microarray (7)
- Human Interferon (6)
- Machine Learning (6)
- Omics (6)
- Omics-LHV Project (6)
- Type 1 Diabetes (6)
- Autoimmunity (5)
- Synthetic (5)
- Biomarkers (4)
- Molecular Profiling (4)
- Processed Data (4)
- Proteomics (4)
The Human Islet Research Network (HIRN) is a large consortia with many research projects focused on understanding how beta cells are lost in type 1 diabetics (T1D) with a goal of finding how to protect against or replace the loss of functional beta cells. The consortia has multiple branches of...
Datasets
0
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson Omics-LHV Profiling of Host Response to Ebola Virus Infection Background Ebola virus ( EBOV ) is a high risk biological agent, belonging to the Flaviviridae family, and is classified as a Category A priority pathogen by the National Institute...
Category
Datasets
6
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson Omics-LHV Profiling of Host Response to Influenza A Virus Infection Background Influenza A virus ( IAV ) is a high risk biological agent belonging to the Orthomyxoviridae family is classified as a Category C priority pathogen by the National...
Category
Datasets
8
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson Omics-LHV Profiling of Host Response to West Nile Virus Infection Background West Nile virus ( WNV ) belongs to the mosquito-borne Flaviviridae family and is classified as a Category A priority pathogen by the National Institute of Allergy and...
Category
Datasets
11
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson Omics-LHV Profiling of Host Response to MERS-CoV Virus Infection Background Middle East Respiratory Syndrome coronavirus ( MERS-CoV ), part of the Coronaviridae family, is classified as a Category C priority pathogen by the National Institute...
Category
Datasets
15
Machine learning is a core technology that is rapidly advancing within type 1 diabetes (T1D) research. Our Human Islet Research Network (HIRN) grant is studying early cellular response initiating β cell stress in T1D through the generation of heterogenous low- and high-throughput molecular...
Datasets
3
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson PNNL DataHub NIAID Program Project: Modeling Host Responses to Understand Severe Human Virus Infections, Multi-Omic Viral Dataset Catalog Collection Background The National Institute of Allergy and Infectious Diseases (NIAID) "Modeling Host...
Category
Datasets
45
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson MERS-CoV Experiment MHAE001 The purpose of this experiment was to evaluate the human host response to wild-type MERS-CoV (icMERS-CoV) virus infection. Sample data was obtained from primary human airway epithelial (HAE) cells for mRNA, miRNA...
Category
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson MERS-CoV Experiment MHAE002 The purpose of this experiment was to evaluate the human host response to wild-type MERS-CoV (strain EMC-2012) virus infection. Sample data was obtained from primary human microvascular endothelial cells (HMVE) for...
Category
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson MERS-CoV Experiment MMVE002 The purpose of this experiment was to evaluate the human host response to MERS-CoV (strain EMC-2012) infectious clone (icMERS-CoV) virus infection. Sample data was obtained from primary human fibroblast cells for...
Category
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson MERS-CoV Experiment MHAE003 The purpose of this experiment was to evaluate the human host response to wild-type MERS-CoV (strain EMC-2012) infection. Sample data was obtained from primary human airway epithelial cells (HAE) for mRNA...
Category
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson MERS-CoV Experiment MM001 The purpose of this experiment was to evaluate the host response to wild-type MERS-CoV virus infection. Sample data was obtained from primary mouse (strain C57BL/6J) whole lung for mRNA, proteomics, metabolomics, and...
Category
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson MERS-CoV Experiment MCL002 The purpose of this experiment was to evaluate the human host response to wild-type Middle Eastern Respiratory Syndrome coronavirus (MERS-CoV) and mutants icMERS-RFP, icMERS-DNSP16, and icMERS-d4B virus infection...
Category
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson MERS-CoV Experiment MFB001 The purpose of this experiment was to evaluate the human host response to wild-type MERS-CoV (icMERS-CoV) virus infection. Sample data was obtained from primary human fibroblasts and processed for mRNA, miRNA...