The srpAnalytics modeling pipeline for the Superfund Research Program Analytics Portal.
Filter results
Category
- (-) Biology (21)
- (-) Computing & Analytics (4)
- Scientific Discovery (31)
- Human Health (12)
- Earth System Science (11)
- Computational Research (7)
- Integrative Omics (7)
- National Security (5)
- Data Analytics & Machine Learning (4)
- Microbiome Science (4)
- Energy Resiliency (2)
- Chemical & Biological Signatures Science (1)
- Chemistry (1)
- Coastal Science (1)
- Computational Mathematics & Statistics (1)
- Energy Efficiency (1)
- Energy Storage (1)
- Renewable Energy (1)
- Solar Energy (1)
- Weapons of Mass Effect (1)
Tags
- Omics-LHV Project (6)
- Virology (6)
- Differential Expression Analysis (5)
- Gene expression profile data (5)
- Immune Response (5)
- Multi-Omics (5)
- Time Sampled Measurement Datasets (5)
- Autoimmunity (4)
- Biomarkers (4)
- Homo sapiens (4)
- Mass spectrometry data (4)
- Molecular Profiling (4)
- Synthetic (4)
- Type 1 Diabetes (4)
- Machine Learning (3)
- Mass spectrometry-based Omics (3)
- Mus musculus (3)
- Omics (3)
- Biological and Environmental Research (2)
- Ebola (2)
- Human Interferon (2)
- Influenza A (2)
- Mass Spectrometry (2)
- MERS-CoV (2)
- Predictive Modeling (2)
- West Nile virus (2)
- DOE (1)
- Genomics (1)
- Output Databases (1)
- Proteomics (1)
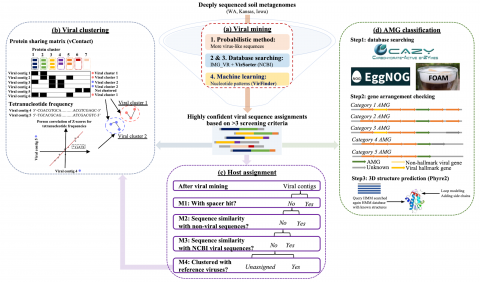
Code pertaining to the Soil Microbiome SFA Project publication data visualizations 'DNA viral diversity, abundance and functional potential vary across grassland soils with a range of historical moisture regimes' for processing publication data downloads.
Category
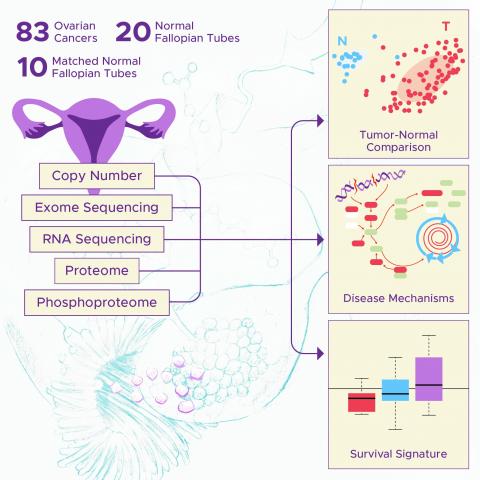
The Human Islet Research Network (HIRN) is a large consortia with many research projects focused on understanding how beta cells are lost in type 1 diabetics (T1D) with a goal of finding how to protect against or replace the loss of functional beta cells. The consortia has multiple branches of...
Datasets
0
This data is a model of synthetic adversarial activity surrounded by noise and was funded by DARPA. The various versions include gradually more complex networks of activities.
Category
Datasets
1
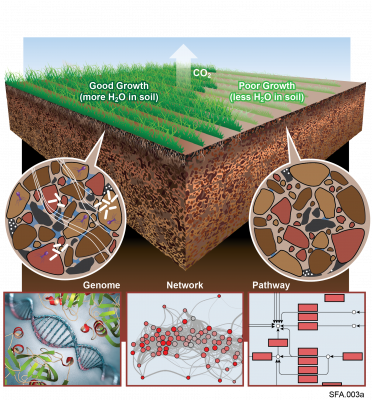
The Phenotypic Response of the Soil Microbiome to Environmental Perturbations Project (Soil Microbiome SFA) at Pacific Northwest National Laboratory is a Genomic Sciences Program Science Focus Area (SFA) Project operating under the Environmental Microbiome Science Research Area. The Soil Microbiome...
Datasets
23
The following R source code was used for plotting figures of the viral communities detected from three grasslands soil metagenomes with a historical precipitation gradient ( WA-TmG.2.0 , KS-TmG.2.0 , IA-TmG.2.0 ) from project publication 'DNA viral diversity, abundance and functional potential vary...
Category
Fusarium sp. DS682 Proteogenomics Statistical Data Analysis of SFA dataset download: 10.25584/KSOmicsFspDS682/1766303 . GitHub Repository Source: https://github.com/lmbramer/Fusarium-sp.-DS-682-Proteogenomics MaxQuant Export Files (txt) Trelliscope Boxplots (jsonp) Fusarium Report (.Rmd, html)...
This data is a model of synthetic adversarial activity surrounded by noise and was funded by DARPA. The various versions include gradually more complex networks of activities.
Category
Datasets
1
Category
Datasets
7
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson Omics-LHV Profiling of Host Interferon-Stimulated Response to Virus Infection Background The human host Interferon ( IFN ) alpha, beta, and gamma participate in the body's natural immune response to lethal virus infection and disease. The...
Category
Datasets
5
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson Omics-LHV Profiling of Host Response to MERS-CoV Virus Infection Background Middle East Respiratory Syndrome coronavirus ( MERS-CoV ), part of the Coronaviridae family, is classified as a Category C priority pathogen by the National Institute...
Category
Datasets
15
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson Omics-LHV Profiling of Host Response to Ebola Virus Infection Background Ebola virus ( EBOV ) is a high risk biological agent, belonging to the Flaviviridae family, and is classified as a Category A priority pathogen by the National Institute...
Category
Datasets
6
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson Omics-LHV Profiling of Host Response to Influenza A Virus Infection Background Influenza A virus ( IAV ) is a high risk biological agent belonging to the Orthomyxoviridae family is classified as a Category C priority pathogen by the National...
Category
Datasets
8
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson Omics-LHV Profiling of Host Response to West Nile Virus Infection Background West Nile virus ( WNV ) belongs to the mosquito-borne Flaviviridae family and is classified as a Category A priority pathogen by the National Institute of Allergy and...
Category
Datasets
11
Category
Datasets
1