Last updated on 2023-01-30T00:09:57+00:00 by LN Anderson Ebola Virus Experiment EIHH001 This experiment evaluated Immortalized Human Hepatocyte ( IHH ) cell line host response to wild-type Ebola viruses Zaire Ebola (ZEBOV '76) and Reston Ebola (REBOV '08) viral infection. Processed Transcriptome...
Filter results
Category
- (-) Human Health (13)
- Scientific Discovery (17)
- Biology (16)
- Integrative Omics (9)
- Computational Research (6)
- Data Analytics & Machine Learning (5)
- Computational Mathematics & Statistics (2)
- Microbiome Science (2)
- Computational Mathematics & Statistics (1)
- Computing & Analytics (1)
- Data Analytics & Machine Learning (1)
- Earth System Science (1)
- National Security (1)
Content type
Tags
- (-) Resource Metadata (8)
- (-) Machine Learning (5)
- Virology (79)
- Immune Response (53)
- Time Sampled Measurement Datasets (50)
- Gene expression profile data (47)
- Differential Expression Analysis (46)
- Homo sapiens (34)
- Mass spectrometry data (30)
- Multi-Omics (28)
- Health (23)
- Virus (23)
- Viruses (23)
- MERS-CoV (18)
- Mus musculus (18)
- West Nile virus (13)
- Ebola (11)
- Influenza A (11)
- Microarray (7)
- Human Interferon (6)
- Omics (6)
- Omics-LHV Project (6)
- Type 1 Diabetes (6)
- Autoimmunity (5)
- Biomarkers (4)
- Molecular Profiling (4)
- Processed Data (4)
- Proteomics (4)
- Genomics (3)
- Sequencer System (3)
Last updated on 2023-01-30T00:09:57+00:00 by LN Anderson Ebola Virus Experiment EIHH002 This experiment evaluated Immortalized Human Hepatocyte ( IHH ) cell line host response to response to genetically-reconstructed Zaire Ebola virus infection. Processed Transcriptome Data Unavailable
Category
MERS-CoV Experiment MDC001 Processed Omics Data Unavailable This experiment evaluated primary human dendritic cells infected with a wild type MERS-CoV (icMERS) virus. Related Experimental Data BioProject: PRJNA315103 GEO: GSE79172 (mRNA transcriptome response) Acknowledgment of Federal Funding The...
Category
Influenza A Experiment IM104 Processed Omics Data Unavailable This Influenza experiment evaluated mouse lung expression after etoposide treatment and infection with a pandemic H1N1 influenza strain. Related Experimental Data BioProject: PRJNA382278 GEO: GSE97555 (mRNA transcriptome response)...
Category
The Environmental Determinants of Diabetes in the Young (TEDDY) study is searching for factors influencing the development of type 1 diabetes (T1D) in children. Research has shown that there are certain genes that correlate to higher risk of developing T1D, but not all children with these genes...
Datasets
1
The Diabetes Autoimmunity Study in the Young (DAISY) seeks to find environmental factors that can trigger the development of type 1 diabetes (T1D) in children. DAISY follows children with high-risk of developing T1D based on family history or genetic markers. Genes, diets, infections, and...
Datasets
1
Machine learning is a core technology that is rapidly advancing within type 1 diabetes (T1D) research. Our Human Islet Research Network (HIRN) grant is studying early cellular response initiating β cell stress in T1D through the generation of heterogenous low- and high-throughput molecular...
Datasets
3
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson Influenza A Virus Experiment ICL105 Metadata The purpose of this experiment was to evaluate the host epigenetic response to Influenza A virus (subtype H5N1) infection. Sample data was obtained from human lung adenocarcinoma cells (Calu-3) and...
Category
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson Influenza A Virus Experiment ICL106 Metadata The purpose of this experiment was to evaluate the human host epigenetic response to Influenza A virus (subtype H5N1) infection. Samples were obtained from human lung adenocarcinoma cell line (Calu...
Category
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson MERS-CoV Experiment MCL004 Metadata The purpose of this experiment was to evaluate the host response to MERS-CoV virus infection. Sample data was obtained from human lung adenocarcinoma cells (Calu-3) and processed for genome binding/occupancy...
Category
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson MERS-CoV Experiment MCL005 Metadata The purpose of this experiment was to evaluate the human host epigenetic response to MERS-CoV virus infection. Samples were obtained from human lung adenocarcinoma cells (Calu-3) and processed for...
Category
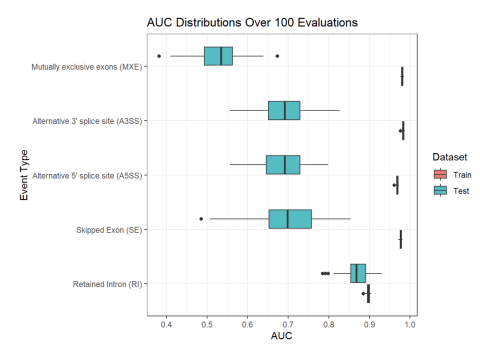
Inclusion levels of alternative splicing (AS) events of five different varieties (i.e. skipped exon (SE), retained intron (RI), alternative 5’ splice site (A5SS), alternative 3’ splice site (A3SS), and mutually exclusive exons (MXE)) were measured in human blood samples from two separate cohorts of...
Comprised of 6,426 sample runs, The Environmental Determinants of Diabetes in the Young (TEDDY) proteomics validation study constitutes one of the largest targeted proteomics studies in the literature to date. Making quality control (QC) and donor sample data available to researchers aligns with...