Last updated on 2023-05-02T18:08:23+00:00 by LN Anderson Fungal Monoisolate Multi-Omics Data Package DOI "KS4A-Omics1.0_FspDS68" Molecular mechanisms underlying fungal mineral weathering and nutrient translocation in low nutrient environments remain poorly resolved, due to the lack of a platform for...
Filter results
Category
- (-) Integrative Omics (74)
- (-) Data Analytics & Machine Learning (9)
- (-) Computational Mathematics & Statistics (1)
- (-) Grid Analytics (1)
- (-) Wind Energy (1)
- Scientific Discovery (309)
- Biology (200)
- Earth System Science (136)
- Human Health (104)
- Microbiome Science (42)
- Computational Research (25)
- National Security (22)
- Computing & Analytics (15)
- Chemistry (10)
- Energy Resiliency (10)
- Computational Mathematics & Statistics (7)
- Materials Science (7)
- Visual Analytics (6)
- Chemical & Biological Signatures Science (5)
- Renewable Energy (5)
- Weapons of Mass Effect (5)
- Atmospheric Science (4)
- Coastal Science (4)
- Data Analytics & Machine Learning (4)
- Ecosystem Science (4)
- Plant Science (3)
- Cybersecurity (2)
- Distribution (2)
- Electric Grid Modernization (2)
- Energy Efficiency (2)
- Energy Storage (2)
- Grid Cybersecurity (2)
- Solar Energy (2)
- Bioenergy Technologies (1)
- High-Performance Computing (1)
- Subsurface Science (1)
- Terrestrial Aquatics (1)
- Transportation (1)
Tags
- Virology (57)
- Immune Response (53)
- Time Sampled Measurement Datasets (50)
- Gene expression profile data (47)
- Differential Expression Analysis (46)
- Homo sapiens (34)
- Mass spectrometry data (30)
- Multi-Omics (30)
- MERS-CoV (18)
- Mus musculus (18)
- West Nile virus (13)
- Ebola (11)
- Influenza A (11)
- Omics (9)
- Resource Metadata (9)
- Mass Spectrometry (7)
- Human Interferon (6)
- Machine Learning (6)
- Microarray (6)
- Omics-LHV Project (6)
- Type 1 Diabetes (6)
- Autoimmunity (5)
- Proteomics (5)
- Biomarkers (4)
- Mass Spectrometer (4)
- Mass spectrometry-based Omics (4)
- Molecular Profiling (4)
- Predictive Modeling (4)
- Imaging (3)
- Microscopy (3)
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson Influenza A Virus Experiment ICL105 Metadata The purpose of this experiment was to evaluate the host epigenetic response to Influenza A virus (subtype H5N1) infection. Sample data was obtained from human lung adenocarcinoma cells (Calu-3) and...
Category
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson Influenza A Virus Experiment ICL106 Metadata The purpose of this experiment was to evaluate the human host epigenetic response to Influenza A virus (subtype H5N1) infection. Samples were obtained from human lung adenocarcinoma cell line (Calu...
Category
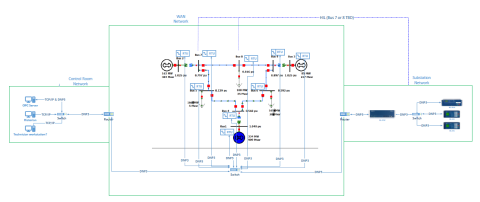
This dataset includes one baseline and three cybersecurity based scenarios utilizing the IEEE 9 Bus Model. This instantiation of the IEEE 9 model was built utilizing the OpalRT Simulator ePhasorsim module, with Bus 7 represented by hardware in the loop (HiL). The HiL was represented by two SEL351s...
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson MERS-CoV Experiment MCL004 Metadata The purpose of this experiment was to evaluate the host response to MERS-CoV virus infection. Sample data was obtained from human lung adenocarcinoma cells (Calu-3) and processed for genome binding/occupancy...
Category
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson MERS-CoV Experiment MCL005 Metadata The purpose of this experiment was to evaluate the human host epigenetic response to MERS-CoV virus infection. Samples were obtained from human lung adenocarcinoma cells (Calu-3) and processed for...
Category
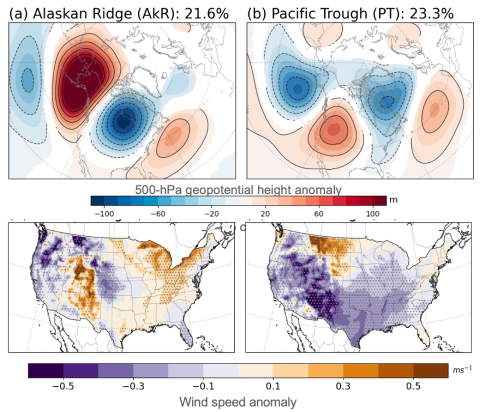
Supporting data and code uploaded to DataHub for "How do the weather regimes drive wind speed and power production at the sub-seasonal to seasonal timescales over the CONUS?" Created by Ye Liu*, Sha Feng*, Yun Qian, Berg K Larry, Huilin Huang *POC: Ye Liu, Ye.Liu@pnnl.gov Sha Feng, sfeng@pnnl.gov --...
Category
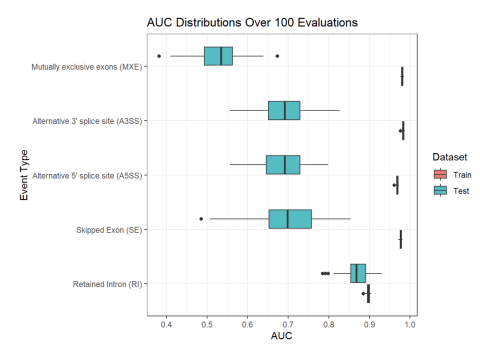
Inclusion levels of alternative splicing (AS) events of five different varieties (i.e. skipped exon (SE), retained intron (RI), alternative 5’ splice site (A5SS), alternative 3’ splice site (A3SS), and mutually exclusive exons (MXE)) were measured in human blood samples from two separate cohorts of...
A total of 172 children from the DAISY study with multiple plasma samples collected over time, with up to 23 years of follow-up, were characterized via proteomics analysis. Of the children there were 40 controls and 132 cases. All 132 cases had measurements across time relative to IA. Sampling was...
Comprised of 6,426 sample runs, The Environmental Determinants of Diabetes in the Young (TEDDY) proteomics validation study constitutes one of the largest targeted proteomics studies in the literature to date. Making quality control (QC) and donor sample data available to researchers aligns with...