"Visualizing the Hidden Half: Plant-Microbe Interactions in the Rhizosphere" Plant roots and the associated rhizosphere constitute a dynamic environment that fosters numerous intra- and interkingdom interactions, including metabolite exchange between plants and soil mediated by root exudates and the...
Filter results
Category
- (-) Microbiome Science (8)
- (-) High-Performance Computing (1)
- Scientific Discovery (38)
- Biology (36)
- Earth System Science (26)
- Integrative Omics (7)
- Chemistry (3)
- Computational Research (3)
- Human Health (2)
- Computational Mathematics & Statistics (1)
- Computing & Analytics (1)
- Data Analytics & Machine Learning (1)
- Ecosystem Science (1)
- National Security (1)
- Plant Science (1)
Content type
Tags
- (-) Mass Spectrometry (3)
- (-) Soil Microbiology (2)
- (-) Whole Genome Sequencing (2)
- (-) Data Analysis (1)
- (-) High-Performance Computing (1)
- Omics (10)
- PerCon SFA (9)
- High Throughput Sequencing (8)
- Genomics (7)
- Sequencer System (5)
- Synthetic Biology (5)
- A. pittii SO1 (2)
- Amplicon Sequencing (2)
- Biological and Environmental Research (2)
- Imaging (2)
- Long Read Sequencer (2)
- Mass spectrometry data (2)
- Proteomics (2)
- RNA Sequence Analysis (2)
- Sorghum bicolor (2)
- Spectroscopy (2)
- Statistical Expression Analysis (2)
- DOE (1)
- Lipidomics (1)
- Machine Learning (1)
- Mass Spectrometer (1)
- metabolomics (1)
- Metagenomics (1)
- Output Databases (1)
- Viruses (1)
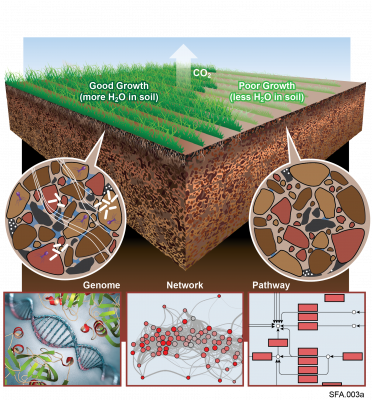
Pending Review Microbiomes contribute to multiple ecosystem services by transforming organic matter in soil. Extreme shifts in the environment, such as drying-rewetting cycles during drought, can impact microbial metabolism of organic matter by altering their physiology and function. These...
Metabolite exchange between plant roots and their associated rhizosphere microbiomes underpins plant growth promotion by microbes. Sorghum bicolor is a cereal crop that feeds animals and humans and is used for bioethanol production. Its root tips exude large amounts of a lipophilic benzoquinone...
"Moisture modulates soil reservoirs of active DNA and RNA viruses" Soil is known to harbor viruses, but the majority are uncharacterized and their responses to environmental changes are unknown. Here, we used a multi-omics approach (metagenomics, metatranscriptomics and metaproteomics) to detect...
The Phenotypic Response of the Soil Microbiome to Environmental Perturbations Project (Soil Microbiome SFA) at Pacific Northwest National Laboratory is a Genomic Sciences Program Science Focus Area (SFA) Project operating under the Environmental Microbiome Science Research Area. The Soil Microbiome...
Datasets
23
Fusarium sp. DS682 Proteogenomics Statistical Data Analysis of SFA dataset download: 10.25584/KSOmicsFspDS682/1766303 . GitHub Repository Source: https://github.com/lmbramer/Fusarium-sp.-DS-682-Proteogenomics MaxQuant Export Files (txt) Trelliscope Boxplots (jsonp) Fusarium Report (.Rmd, html)...
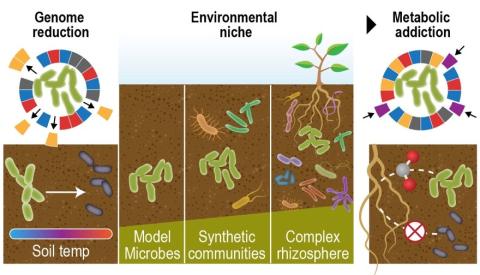
Last updated on 2023-02-23T19:37:46+00:00 by LN Anderson PerCon SFA Project Publication Experimental Data Catalog The Persistence Control of Engineered Functions in Complex Soil Microbiomes Project (PerCon SFA) at Pacific Northwest National Laboratory ( PNNL ) is a Genomic Sciences Program...
Datasets
3
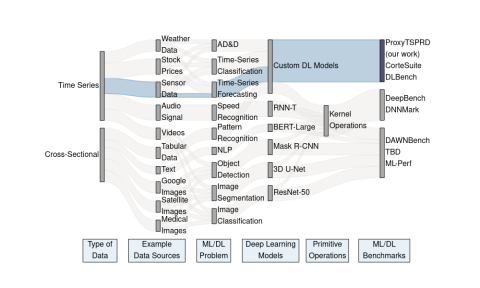
ProxyTSPRD profiles are collected using NVIDIA Nsight Systems version 2020.3.2.6-87e152c and capture computational patterns from training deep learning-based time-series proxy-applications on four different levels: models (Long short-term Memory and Convolutional Neural Network), DL frameworks...
Last updated on 2023-05-31T16:35:53+00:00 by LN Anderson PerCon SFA: Sequencing of Sorgoleone Promoting Rhizobacteria Isolates Whole genome sequencing (WGS) of sorgoleone utilizing rhizobacteria strains Pseudomonas sorgoleonovorans SO81 , Burkholderia anthina SO82 , and Acinetobacter pittii SO1 , as...