Please cite as : Anderson L.N., R. Wu, W.C. Nelson, J.E. McDermott, K.S. Hofmockel, and J.K. Jansson. 2021. WA-TmG.2.0 (Metagenome, WA). [Data Set] PNNL DataHub. https://doi.org/10.25584/WATmG2/1770324 Soil samples were collected in triplicate in the fall of 2017 across the three grassland...
Filter results
Content type
Dataset Type
- (-) Metabolomics (28)
- (-) Metagenomics (9)
- Omics (86)
- Transcriptomics (51)
- Proteomics (39)
- Sequencing (39)
- Lipidomics (26)
- Simulation Results (17)
- Amplicon (16S, ITS) (15)
- Microarray (9)
- Genomics (4)
- Computed Analysis (3)
- ChIP-seq (2)
- Geospacial (2)
- Staff (2)
- Cybersecurity (1)
- Electromagnetic Spectrum (1)
- Mass Spectra (1)
- Microscopy (1)
- Phenomics (1)
- Whole genome sequencing (WGS) (1)
Tags
- Virology (26)
- Differential Expression Analysis (23)
- Immune Response (23)
- Mass spectrometry data (23)
- Time Sampled Measurement Datasets (23)
- Multi-Omics (22)
- Gene expression profile data (17)
- Homo sapiens (16)
- MERS-CoV (12)
- Mus musculus (7)
- Soil Microbiology (7)
- Metagenomics (6)
- sequencing (6)
- Influenza A (5)
- West Nile virus (4)
- Health (3)
- metagenomics (3)
- Microbiome (3)
- Omics (3)
- Sequencing (3)
- soil microbiology (3)
- Virus (3)
- Viruses (3)
- Ebola (2)
- IAREC (2)
- omics (2)
- Amplicon Sequencing (1)
- Mass Spectrometry (1)
- MPLEX (1)
- Software Data Analysis (1)
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson West Nile Virus Experiment WLN003 The purpose of this experiment was to evaluate the host response to West Nile virus (strain WNV-NY99) wild-type clone 382 and mutant 382-E218A 2 nt virus infection. Sample data was obtained from mouse (strain...
Category
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson West Nile Virus Experiment WSE001 The purpose of this experiment was to evaluate the host response to West Nile virus (strain WNV-NY99) wild-type clone 382 virus infection. Sample data was obtained from mouse (strain C57BL6/JAX) blood serum...
Category
Please cite as : Anderson L.N., R. Wu, W.C. Nelson, J.E. McDermott, K.S. Hofmockel, and J.K. Jansson. 2021. KS-TmG.2.0 (Metagenome, KS). [Data Set] PNNL DataHub. https://doi.org/10.25584/KSTmG2/1770332 Soil samples were collected in triplicate in the fall of 2017 across the three grassland locations...
Category
Please cite as : Anderson L.N., R. Wu, W.C. Nelson, J.E. McDermott, K.S. Hofmockel, and J.K. Jansson. 2021. IA-TmG.2.0 (Metagenome, IA). [Data Set] PNNL DataHub. https://doi.org/10.25584/IATmG2/1770333 Soil samples were collected in triplicate in the fall of 2017 across the three grassland locations...
Category
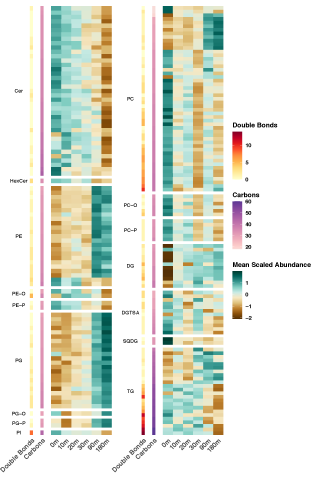
Rapid remodeling of the soil lipidome in response to a drying-rewetting event - Multi-Omics Data Package DOI Data package contents reported here are the first version and contain pre- and post-processed data acquisition and subsequent downstream analysis files using various data source instrument...
Category
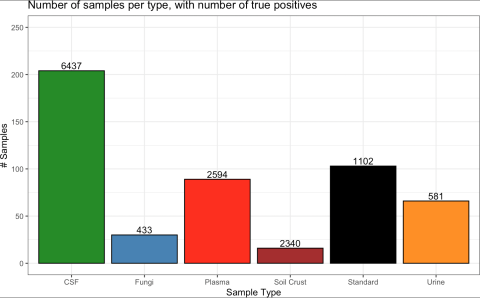
Data comprised of 204, 103, 89, 66, 30, and 16 gas chromatography mass spectrometry (GC-MS) runs from human cerebrospinal fluid (CSF), standards, human blood plasma, human urine, fungi species (A. niger, A. nidulans, T. reesei), and soil crust, respectively Across 4,523,251 candidate matches, 13,487...