Last updated on 2023-01-30T00:09:57+00:00 by LN Anderson Ebola Virus Experiment EIHH001 This experiment evaluated Immortalized Human Hepatocyte ( IHH ) cell line host response to wild-type Ebola viruses Zaire Ebola (ZEBOV '76) and Reston Ebola (REBOV '08) viral infection. Processed Transcriptome...
Filter results
Category
- Biology (71)
- Scientific Discovery (71)
- Human Health (59)
- Integrative Omics (55)
- Earth System Science (10)
- Computational Research (6)
- Data Analytics & Machine Learning (5)
- Microbiome Science (4)
- Computational Mathematics & Statistics (2)
- Computational Mathematics & Statistics (1)
- Computing & Analytics (1)
- Data Analytics & Machine Learning (1)
- Materials Science (1)
- National Security (1)
- Plant Science (1)
Tags
- (-) Immune Response (53)
- (-) Influenza A (11)
- (-) Machine Learning (7)
- Virology (79)
- Time Sampled Measurement Datasets (50)
- Gene expression profile data (47)
- Differential Expression Analysis (46)
- Homo sapiens (34)
- Mass spectrometry data (31)
- Multi-Omics (30)
- Viruses (26)
- Omics (25)
- Health (23)
- Virus (23)
- Soil Microbiology (21)
- MERS-CoV (18)
- Mus musculus (18)
- Mass Spectrometry (14)
- Synthetic (14)
- sequencing (13)
- West Nile virus (13)
- Genomics (12)
- Ebola (11)
- PerCon SFA (10)
- High Throughput Sequencing (9)
- Metagenomics (9)
- Resource Metadata (9)
- Microbiome (8)
- Proteomics (8)
- Microarray (7)
Last updated on 2023-01-30T00:09:57+00:00 by LN Anderson Ebola Virus Experiment EIHH002 This experiment evaluated Immortalized Human Hepatocyte ( IHH ) cell line host response to response to genetically-reconstructed Zaire Ebola virus infection. Processed Transcriptome Data Unavailable
Category
Category
Category
Influenza A Experiment IM104 Processed Omics Data Unavailable This Influenza experiment evaluated mouse lung expression after etoposide treatment and infection with a pandemic H1N1 influenza strain. Related Experimental Data BioProject: PRJNA382278 GEO: GSE97555 (mRNA transcriptome response)...
Category
Soil fungi facilitate the translocation of inorganic nutrients from soil minerals to other microorganisms and plants. This ability is particularly advantageous in impoverished soils, because fungal mycelial networks can bridge otherwise spatially disconnected and inaccessible nutrient hotspots...
Category
"DNA Viral Diversity, Abundance, and Functional Potential Vary across Grassland Soils with a Range of Historical Moisture Regimes" Soil viruses are abundant, but the influence of the environment and climate on soil viruses remains poorly understood. Here, we addressed this gap by comparing the...
Category
"Visualizing the Hidden Half: Plant-Microbe Interactions in the Rhizosphere" Plant roots and the associated rhizosphere constitute a dynamic environment that fosters numerous intra- and interkingdom interactions, including metabolite exchange between plants and soil mediated by root exudates and the...
The recently developed real-time nuclear–electronic orbital (RT-NEO) approach provides an elegant framework for treating electrons and selected nuclei, typically protons, quantum mechanically in nonequilibrium dynamical processes. However, the RT-NEO approach neglects the motion of the other nuclei...
A major challenge in biotechnology and biomanufacturing is the identification of a set of biomarkers for perturbations and metabolites of interest. Here, we develop a data-driven, transcriptome-wide approach to rank perturbation-inducible genes from time-series RNA sequencing data for the discovery...
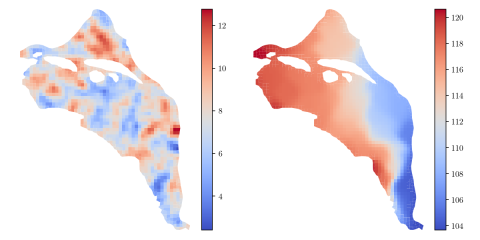
HDF5 file containing 10,000 hydraulic transmissivity inputs and the corresponding hydraulic pressure field outputs for a two-dimensional saturated flow model of the Hanford Site. The inputs are generated by sampling a 1,000-dimensional Kosambi-Karhunen-Loève (KKL) model of the transmissivity field...
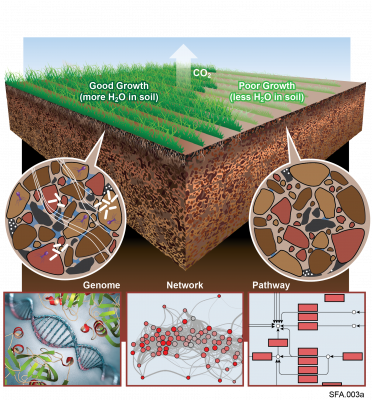
The Phenotypic Response of the Soil Microbiome to Environmental Perturbations Project (Soil Microbiome SFA) at Pacific Northwest National Laboratory is a Genomic Sciences Program Science Focus Area (SFA) Project operating under the Environmental Microbiome Science Research Area. The Soil Microbiome...
Datasets
23
Fusarium sp. DS682 Proteogenomics Statistical Data Analysis of SFA dataset download: 10.25584/KSOmicsFspDS682/1766303 . GitHub Repository Source: https://github.com/lmbramer/Fusarium-sp.-DS-682-Proteogenomics MaxQuant Export Files (txt) Trelliscope Boxplots (jsonp) Fusarium Report (.Rmd, html)...
The Environmental Determinants of Diabetes in the Young (TEDDY) study is searching for factors influencing the development of type 1 diabetes (T1D) in children. Research has shown that there are certain genes that correlate to higher risk of developing T1D, but not all children with these genes...
Datasets
1
The Diabetes Autoimmunity Study in the Young (DAISY) seeks to find environmental factors that can trigger the development of type 1 diabetes (T1D) in children. DAISY follows children with high-risk of developing T1D based on family history or genetic markers. Genes, diets, infections, and...
Datasets
1