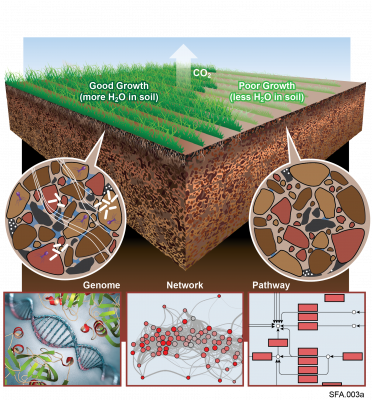
"Visualizing the Hidden Half: Plant-Microbe Interactions in the Rhizosphere" Plant roots and the associated rhizosphere constitute a dynamic environment that fosters numerous intra- and interkingdom interactions, including metabolite exchange between plants and soil mediated by root exudates and the...
Filter results
Category
- (-) Microbiome Science (10)
- (-) Integrative Omics (8)
- (-) Data Analytics & Machine Learning (5)
- Scientific Discovery (72)
- Biology (49)
- Earth System Science (28)
- Human Health (25)
- Computational Research (11)
- National Security (7)
- Computing & Analytics (5)
- Materials Science (5)
- Energy Resiliency (3)
- Chemical & Biological Signatures Science (2)
- Chemistry (2)
- Computational Mathematics & Statistics (2)
- Renewable Energy (2)
- Weapons of Mass Effect (2)
- Atmospheric Science (1)
- Coastal Science (1)
- Data Analytics & Machine Learning (1)
- Energy Efficiency (1)
- Energy Storage (1)
- Plant Science (1)
- Solar Energy (1)
Tags
- Omics-LHV Project (6)
- Virology (6)
- Differential Expression Analysis (5)
- Gene expression profile data (5)
- Immune Response (5)
- Multi-Omics (5)
- PerCon SFA (5)
- Time Sampled Measurement Datasets (5)
- Autoimmunity (4)
- Biomarkers (4)
- Homo sapiens (4)
- Machine Learning (4)
- Mass spectrometry data (4)
- Molecular Profiling (4)
- Omics (4)
- Synthetic Biology (4)
- Type 1 Diabetes (4)
- High Throughput Sequencing (3)
- Mass Spectrometry (3)
- Mass spectrometry-based Omics (3)
- Mus musculus (3)
- Biological and Environmental Research (2)
- Ebola (2)
- Human Interferon (2)
- Imaging (2)
- Influenza A (2)
- MERS-CoV (2)
- RNA Sequence Analysis (2)
- Spectroscopy (2)
- West Nile virus (2)
Pending Review Microbiomes contribute to multiple ecosystem services by transforming organic matter in soil. Extreme shifts in the environment, such as drying-rewetting cycles during drought, can impact microbial metabolism of organic matter by altering their physiology and function. These...
Elmore JR, Dexter GN, Baldino H, Huenemann JD, Francis R, Peabody GL 5th, Martinez-Baird J, Riley LA, Simmons T, Coleman-Derr D, Guss AM, Egbert RG. High-throughput genetic engineering of nonmodel and undomesticated bacteria via iterative site-specific genome integration. Sci Adv. 2023 Mar 10;9(10)...
The rhizosphere represents a dynamic and complex interface between plant hosts and the microbial community found in the surrounding soil. While it is recognized that manipulating the rhizosphere has the potential to improve plant fitness and health, engineering the rhizosphere microbiome through...
Agriculture is the largest source of greenhouse gases (GHG) production. Conversion of nitrogen fertilizers into more reduced forms by microbes through a process known as biological nitrification drives GHG production, enhances proliferation of toxic algal blooms, and increases cost of crop...
Metabolite exchange between plant roots and their associated rhizosphere microbiomes underpins plant growth promotion by microbes. Sorghum bicolor is a cereal crop that feeds animals and humans and is used for bioethanol production. Its root tips exude large amounts of a lipophilic benzoquinone...
A major challenge in biotechnology and biomanufacturing is the identification of a set of biomarkers for perturbations and metabolites of interest. Here, we develop a data-driven, transcriptome-wide approach to rank perturbation-inducible genes from time-series RNA sequencing data for the discovery...
The Human Islet Research Network (HIRN) is a large consortia with many research projects focused on understanding how beta cells are lost in type 1 diabetics (T1D) with a goal of finding how to protect against or replace the loss of functional beta cells. The consortia has multiple branches of...
Datasets
0
The Phenotypic Response of the Soil Microbiome to Environmental Perturbations Project (Soil Microbiome SFA) at Pacific Northwest National Laboratory is a Genomic Sciences Program Science Focus Area (SFA) Project operating under the Environmental Microbiome Science Research Area. The Soil Microbiome...
Datasets
23
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson Omics-LHV Profiling of Host Response to Influenza A Virus Infection Background Influenza A virus ( IAV ) is a high risk biological agent belonging to the Orthomyxoviridae family is classified as a Category C priority pathogen by the National...
Category
Datasets
8
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson Omics-LHV Profiling of Host Response to West Nile Virus Infection Background West Nile virus ( WNV ) belongs to the mosquito-borne Flaviviridae family and is classified as a Category A priority pathogen by the National Institute of Allergy and...
Category
Datasets
11
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson Omics-LHV Profiling of Host Interferon-Stimulated Response to Virus Infection Background The human host Interferon ( IFN ) alpha, beta, and gamma participate in the body's natural immune response to lethal virus infection and disease. The...
Category
Datasets
5
Last updated on 2024-02-11T22:41:43+00:00 by LN Anderson PNNL DataHub NIAID Program Project: Modeling Host Responses to Understand Severe Human Virus Infections, Multi-Omic Viral Dataset Catalog Collection Background The National Institute of Allergy and Infectious Diseases (NIAID) "Modeling Host...
Category
Datasets
45
Spruce and Peatlands Responses Under Changing Environments (SPRUCE) site is the 8.1-ha S1 bog, a Picea mariana [black spruce] – Sphagnum spp. ombrotrophic bog forest in northern Minnesota, 40 km north of Grand Rapids, in the USDA Forest Service Marcell Experimental Forest (MEF). Two field research...
Category
Datasets
13
Datasets
1