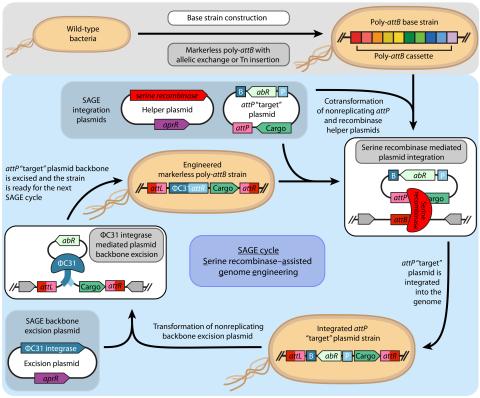
Elmore JR, Dexter GN, Baldino H, Huenemann JD, Francis R, Peabody GL 5th, Martinez-Baird J, Riley LA, Simmons T, Coleman-Derr D, Guss AM, Egbert RG. High-throughput genetic engineering of nonmodel and undomesticated bacteria via iterative site-specific genome integration. Sci Adv. 2023 Mar 10;9(10)...
Filter results
Category
- (-) Microbiome Science (7)
- (-) Computing & Analytics (1)
- Scientific Discovery (19)
- Biology (18)
- Human Health (9)
- Computational Research (6)
- Data Analytics & Machine Learning (5)
- Earth System Science (5)
- Integrative Omics (5)
- National Security (3)
- Chemical & Biological Signatures Science (2)
- Weapons of Mass Effect (2)
- Chemistry (1)
- Computational Mathematics & Statistics (1)
- Computational Mathematics & Statistics (1)
- Data Analytics & Machine Learning (1)
Content type
Tags
- (-) Biological and Environmental Research (2)
- (-) Machine Learning (2)
- (-) Proteomics (2)
- (-) Statistical Expression Analysis (2)
- Omics (9)
- PerCon SFA (9)
- High Throughput Sequencing (8)
- Genomics (7)
- Sequencer System (5)
- Synthetic (5)
- Synthetic Biology (5)
- Mass Spectrometry (3)
- A. pittii SO1 (2)
- Amplicon Sequencing (2)
- Imaging (2)
- Long Read Sequencer (2)
- Mass spectrometry data (2)
- RNA Sequence Analysis (2)
- Sorghum bicolor (2)
- Spectroscopy (2)
- Whole Genome Sequencing (2)
- Data Analysis (1)
- DOE (1)
- Environmental Microbiome Science (1)
- High-Performance Computing (1)
- Lipidomics (1)
- Mass Spectrometer (1)
- metabolomics (1)
- Output Databases (1)
- Soil Microbiology (1)
A major challenge in biotechnology and biomanufacturing is the identification of a set of biomarkers for perturbations and metabolites of interest. Here, we develop a data-driven, transcriptome-wide approach to rank perturbation-inducible genes from time-series RNA sequencing data for the discovery...
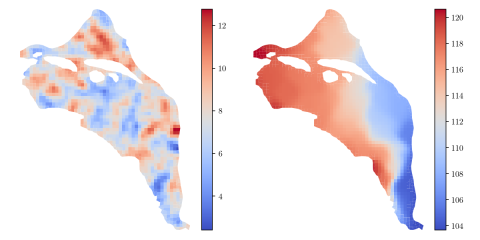
HDF5 file containing 10,000 hydraulic transmissivity inputs and the corresponding hydraulic pressure field outputs for a two-dimensional saturated flow model of the Hanford Site. The inputs are generated by sampling a 1,000-dimensional Kosambi-Karhunen-Loève (KKL) model of the transmissivity field...
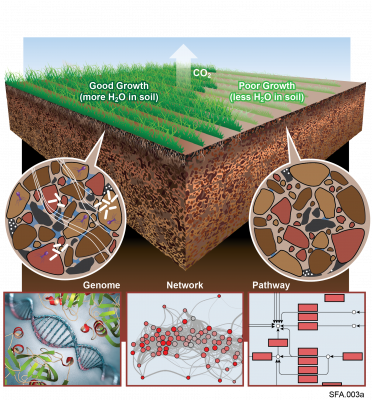
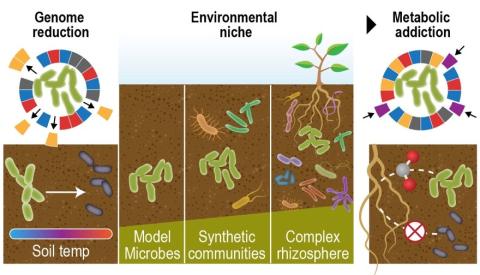
The Phenotypic Response of the Soil Microbiome to Environmental Perturbations Project (Soil Microbiome SFA) at Pacific Northwest National Laboratory is a Genomic Sciences Program Science Focus Area (SFA) Project operating under the Environmental Microbiome Science Research Area. The Soil Microbiome...
Datasets
23
Last updated on 2023-02-23T19:37:46+00:00 by LN Anderson PerCon SFA Project Publication Experimental Data Catalog The Persistence Control of Engineered Functions in Complex Soil Microbiomes Project (PerCon SFA) at Pacific Northwest National Laboratory ( PNNL ) is a Genomic Sciences Program...
Datasets
3
Last updated on 2024-03-03T02:26:52+00:00 by LN Anderson The Thermo Scientific™ Q Exactive™ Plus Mass Spectrometer benchtop LC-MS/MS system combines quadruple precursor ion selection with high-resolution, accurate-mass (HRAM) Orbitrap detection to deliver exceptional performance and versatility...
Last updated on 2023-03-21T18:35:22+00:00 by LN Anderson SAGE-RTP RT-PCR Amplicon Sequencing Barcode Count Analysis Promoter expression data for five bacterial species associated with the Serine recombinase Assisted Genome Engineering (SAGE) research project. Raw Measurement Data NCBI BioProject...
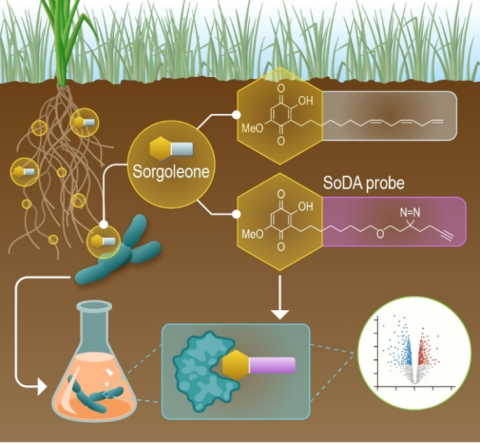
Last updated on 2024-04-19T19:12:08+00:00 by LN Anderson PerCon SFA: Profiling sorghum-microbe interactions with a specialized photoaffinity probe identifies key sorgoleone binders in Acinetobacter pitti Mass spectrometry-based global proteome analysis and SoDA-PAL photoaffinity probe labeled...