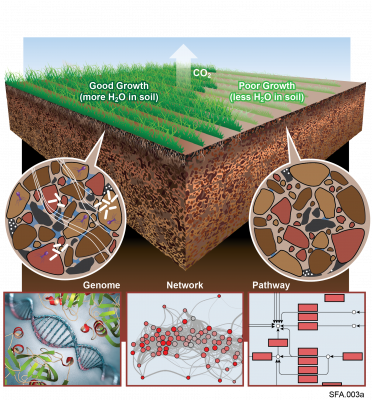
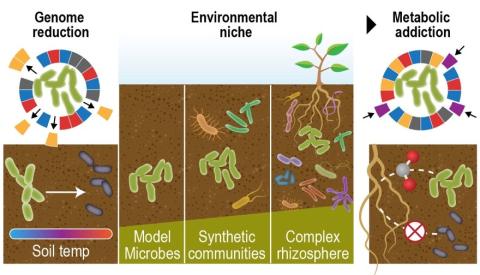
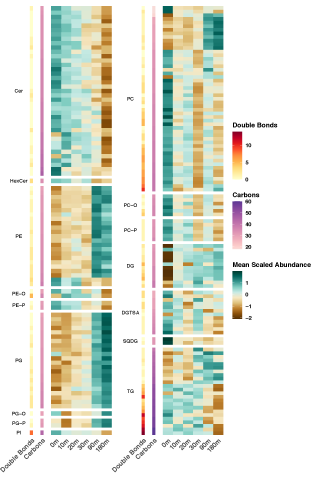
Complete replicate terabase metagenome (TmG.2.0) of grassland soil microbiome collections from KPBS field site in Manhattan, KS. Metagenome (unclassified soil sequencing) Data DOI Package, version 2.0.
Filter results
Tags
- (-) Mass Spectrometry (7)
- (-) Sequencing (4)
- (-) Climate Change (3)
- Omics (22)
- Soil Microbiology (21)
- sequencing (13)
- Genomics (10)
- Metagenomics (9)
- Microbiome (8)
- Fungi (6)
- High Throughput Sequencing (6)
- Imaging (6)
- Type 1 Diabetes (6)
- Autoimmunity (5)
- Machine Learning (5)
- Mass Spectrometer (5)
- Biomarkers (4)
- metagenomics (4)
- Microscopy (4)
- Molecular Profiling (4)
- Sequencer System (4)
- Software Data Analysis (4)
- soil microbiology (4)
- Spectroscopy (4)
- IAREC (3)
- metabolomics (3)
- PerCon SFA (3)
- Proteomics (3)
- Synthetic Biology (3)
- Viruses (3)
Complete replicate terabase metagenome (TmG.2.0) of grassland soil microbiome collections from COBS field site in Boone County, IA. Metagenome (unclassified soil sequencing) Data DOI Package, version 2.0.
Category
Complete replicate terabase metagenome (TmG.2.0) of grassland soil microbiome collections from IAREC field site in Prosser, WA. Metagenome (unclassified soil sequencing) Data DOI Package, version 2.0.
Category
Viral communities detected from three large grassland soil metagenomes with historically different precipitation moisture regimes.
Category
"DNA Viral Diversity, Abundance, and Functional Potential Vary across Grassland Soils with a Range of Historical Moisture Regimes" Soil viruses are abundant, but the influence of the environment and climate on soil viruses remains poorly understood. Here, we addressed this gap by comparing the...
Category
"Visualizing the Hidden Half: Plant-Microbe Interactions in the Rhizosphere" Plant roots and the associated rhizosphere constitute a dynamic environment that fosters numerous intra- and interkingdom interactions, including metabolite exchange between plants and soil mediated by root exudates and the...
pmartR Software Overview The pmartR package provides a single software tool for QC (filtering and normalization), exploratory data analysis (EDA), and statistical analysis (robust to missing data) and includes numerous visualization capabilities of mass spectrometry (MS) omics data (proteomic...
The Phenotypic Response of the Soil Microbiome to Environmental Perturbations Project (Soil Microbiome SFA) at Pacific Northwest National Laboratory is a Genomic Sciences Program Science Focus Area (SFA) Project operating under the Environmental Microbiome Science Research Area. The Soil Microbiome...
Datasets
23
Fusarium sp. DS682 Proteogenomics Statistical Data Analysis of SFA dataset download: 10.25584/KSOmicsFspDS682/1766303 . GitHub Repository Source: https://github.com/lmbramer/Fusarium-sp.-DS-682-Proteogenomics MaxQuant Export Files (txt) Trelliscope Boxplots (jsonp) Fusarium Report (.Rmd, html)...
Category
Datasets
3
Last updated on 2023-02-23T19:37:46+00:00 by LN Anderson PerCon SFA Project Publication Experimental Data Catalog The Persistence Control of Engineered Functions in Complex Soil Microbiomes Project (PerCon SFA) at Pacific Northwest National Laboratory ( PNNL ) is a Genomic Sciences Program...
Datasets
3
The IONTOF TOF.SIMS 5 data source is a time-of-flight secondary ion mass spectrometer and powerful surface analysis tool used to investigate scientific questions in biological, environmental, and energy research. Among the most sensitive of surface analysis tools, it uses a high-vacuum technique...
Category
Category
Rapid remodeling of the soil lipidome in response to a drying-rewetting event - Multi-Omics Data Package DOI Data package contents reported here are the first version and contain pre- and post-processed data acquisition and subsequent downstream analysis files using various data source instrument...